
Advances in Amplification-Free Nucleic Acid Detection via CRISPR-Cas System: From Cascade Circuits to Advanced Readouts
ⓒ The Korean Sensors Society
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License(https://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
While enzymatic amplification remains essential for achieving ultrasensitive detection in molecular diagnostics, amplification-free detection methods offer distinct advantages in simplicity, speed, and cost-effectiveness, particularly for point-of‐care applications. Their value is particularly evident in scenarios where enzymatic amplification is not a viable option. This includes resource-limited settings that lack sophisticated equipment, as well as applications such as live-cell imaging, where amplification is inherently unfeasible. However, the widespread adoption of these techniques has been hindered by limited sensitivity. Recently, clustered regularly interspaced short palindromic repeats (CRISPR)-Cas have been adapted into a powerful platform for molecular diagnostics, providing a promising solution to this sensitivity bottleneck. By leveraging the efficient signal amplification generated by the trans-cleavage activity of Cas12 or Cas13, these systems achieve remarkable sensitivity within a simple, isothermal format. This review summarizes recent advances in amplification-free nucleic acid detection using the CRISPR-Cas system, beginning with a fundamental overview of key Cas proteins and then systematically describing how Cas12 and Cas13 contribute differently to detection, reflecting their unique characteristics. By summarizing these advances, this review aims to provide valuable insights for the future development and application of CRISPR-based amplification-free biosensors.
Keywords:
Nucleic acid detection, CRISPR, Amplification-free, Biosensor1. INTRODUCTION
Nucleic acids, specifically DNA and RNA, are fundamental molecules that carry the genetic information for all living organisms and viruses. The precise detection of specific DNA or RNA sequences has become a cornerstone of modern molecular biology. This capability is essential for various applications, including early diagnosis of infectious diseases [1] and monitoring cancer biomarkers [2]. In this context, the development of rapid, sensitive, and highly specific nucleic acid detection technologies remains a top priority in molecular biology, driving continuous innovation in the field.
For decades, the polymerase chain reaction (PCR) has served as the gold standard in nucleic acid detection [1,3]. The ability to exponentially amplify trace amounts of target sequence provides unparalleled sensitivity and specificity, enabling the detection down to the single-molecule level. However, there are significant hurdles to the broader utility of PCR-based methods. The reliance on sophisticated thermal cyclers, the need for expensive reagents and complex procedures, and the requirement for trained experts confine their use to centralized, well-equipped laboratories. These constraints hinder their deployment for point-of-care testing (POCT) in resource-limited settings [4,5].
To overcome the inherent limitations of PCR, significant research efforts have been reported. The first approach focuses on streamlining the complex PCR procedures within a dedicated device [6,7]. This strategy often leverages recent advances in photothermal PCR, which allows for the miniaturization of thermocyclers. Concurrently, recent advances in microfluidics enable automated workflows, enhancing accessibility for non-specialist operators. Nevertheless, while these integrated systems represent a significant improvement, core challenges remain, such as inherent constraints on further miniaturization and the fundamental reliance on expensive reagents. A second approach involves isothermal amplification technologies, such as loop-mediated isothermal amplification (LAMP) and recombinase polymerase amplification (RPA), which operate at a single, constant temperature [8-12]. By obviating the need for thermal cycling, these methods have enabled the development of simpler, faster, and more portable diagnostic platforms. However, since these approaches still rely on enzymatic amplification, they remain susceptible to non-specific amplification, which can lead to false-positive results and necessitate complex assay optimization.
To address the inherent drawbacks of enzymatic amplification, amplification-free strategies have been developed as a compelling alternative, such as hybridization chain reaction (HCR) [13] and catalytic hairpin assembly (CHA) [14,15]. The primary advantage of these systems lies in their simplicity, which allows integration into portable, cost-effective, instrument-free devices. This simplicity also increases their potential for rapidity and accessibility. Furthermore, a unique advantage of amplification-free strategies is their robust performance in restricted environments where enzymatic amplification is unfeasible, such as in live-cell imaging [16]. Amplification reactions, which often require high temperatures, are fundamentally incompatible with cell viability. Moreover, even for isothermal methods operating at lower temperatures, maintaining precisely optimized conditions is difficult within the complex cellular milieu. Despite these compelling advantages, the foremost challenge for amplification-free approaches is their inherently limited sensitivity, primarily due to the slow kinetics of signal amplification [17]. This limitation creates a clear need for novel strategies that can introduce signal amplification while maintaining the simplicity of the amplification-free approach.
The clustered regularly interspaced short palindromic repeats (CRISPR)-Cas systems, derived from gene-editing technology, have been adapted into a powerful platform for molecular diagnostics [18,19]. These systems offer significant advantages, including a mild reaction temperature, excellent recognition specificity, and high-efficiency signal amplification. Building on these promising features, numerous research groups have developed hybrid strategies that couple the CRISPR-Cas system with other amplification-free technologies, aiming to enhance its performance further and broaden its applicability in diagnostics.
This review provides a detailed overview of recent advances in amplification-free nucleic acid detection, driven by the advent of CRISPR-Cas systems. This review begins with a fundamental overview of the key Cas proteins—including Cas9, Cas12, and Cas13—and elucidates the basic mechanisms of signal amplification. It then delves into how each of these Cas proteins has been ingeniously integrated into amplification-free technologies. This includes strand displacement-based cascade amplification strategies, optimization of CRISPR kinetics, and ultrasensitive readout modalities such as electrochemical, Surface-enhanced Raman scattering (SERS), and digital assays. To conclude, this review offers a comprehensive summary of the current landscape of CRISPR-based amplification-free nucleic acid detection and presents a future perspective in the field.
2. OVERVIEW OF CRISPR-Cas SYSTEMS
To help understand the CRISPR-Cas systems, this section provides an overview of three types of systems: CRISPR-Cas9, CRISPR-Cas12, and CRISPR-Cas13.
2.1 CRISPR-Cas9
Cas9 recognizes and cleaves the specific double-stranded DNA (dsDNA). The working principle of CRISPR-Cas9 is as follows: single-guide RNA (sgRNA)-guided recognition of the protospacer adjacent motif (PAM) site, unwinding of dsDNA, stepwise sgRNA-DNA base-pairing, and cleavage. Cas9 has two active sites, HNH and RuvC, which cleave the complementary and non-complementary strands of the target DNA in a single turnover reaction [20,21]. Since Cas9 lacks signal amplification via collateral cleavage, research has often focused on leveraging its excellent recognition specificity and sensitive readout strategies to detect single-nucleotide mutations [22].
2.2 CRISPR-Cas12a
Similar to Cas9, Cas12a is a single-stranded RNA (ssRNA)-guided DNA endonuclease. Although Cas12a and Cas9 share many functional similarities, Cas12a possesses unique features. Unlike Cas9, Cas12a features a single RuvC nuclease domain. In addition to recognizing dsDNA, CRISPR-Cas12a can recognize single-stranded DNA (ssDNA) [23]. Additionally, Cas12a exhibits both cis-cleavage and trans-cleavage activity [24], while Cas9 exhibits only cis-cleavage activity. The trans-cleavage of CRISPR-Cas12a is a multiple-turnover reaction. This is a key strategy for enhancing detection sensitivity, as it enables signal amplification by repeatedly cleaving the fluorophore-quencher dual-tagged DNA that generates the signal [23].
2.3 CRISPR-Cas13a
Cas13a is an ssRNA-guided endonuclease that recognizes and degrades complementary ssRNA targets [18]. Cas13a recognizes the target ssRNA and induces highly efficient trans-cleavage activity. The difference between Cas12a and Cas13a is the trans-cleavage efficiency. Cas13a has a higher cleavage efficiency than Cas12a, enabling its widespread use in numerous other diagnostic applications [19]. Particularly, among all the Cas13a homologs, LbuCas13a (Cas13a from Leptotrichia buccalis) exhibits superior trans-cleavage activity, where the turnover per single Cas13a-crRNA complex can reach at least 104 [18]. Due to its exceptionally high catalytic efficiency, Cas13a is widely utilized in amplification-free approaches [25].
Table 1 summarizes the features of these three types of CRISPR-Cas systems, unlike Cas9, Cas12a, and Cas13a, which intriguingly exhibit trans-cleavage activities on ssDNA or ssRNA, respectively. Therefore, this review first discusses how the trans-cleavage of Cas12a and Cas13a has advanced amplification-free detection technologies. Subsequently, technologies that directly measure the initial activity of the Cas protein itself based on sensitive readout strategies, without the signal amplification step, are reviewed.
3. CRISPR-Cas12a
3.1 Cas12a-based nucleic acid detection
Among various Cas12a homologs, LbCas12a (Cas12a from Lachnospiraceae bacterium) exhibits superior trans-cleavage activity, where the cleavage activity is 2 orders of magnitude higher than that of AsCas12a [26]. Therefore, amplification-free detection methods have mainly relied on trans-cleavage by LbCas12a to achieve high sensitivity. For instance, He et al. reported amplification-free detection of African swine fever viral DNA using LbCas12a (Fig. 1(a)) [27]. Without target amplification, they achieved a detection limit of 1 pM with 8 hours of CRISPR reaction time. Another study reported that reducing the reaction time to 15 minutes worsened the detection limit to 100 pM [28]. Despite their simplicity, these detection performances reveal a significant gap compared to the sensitivity and rapidity required for infectious disease diagnosis. Therefore, strategies to enhance the signal from trans-cleavage are necessary.
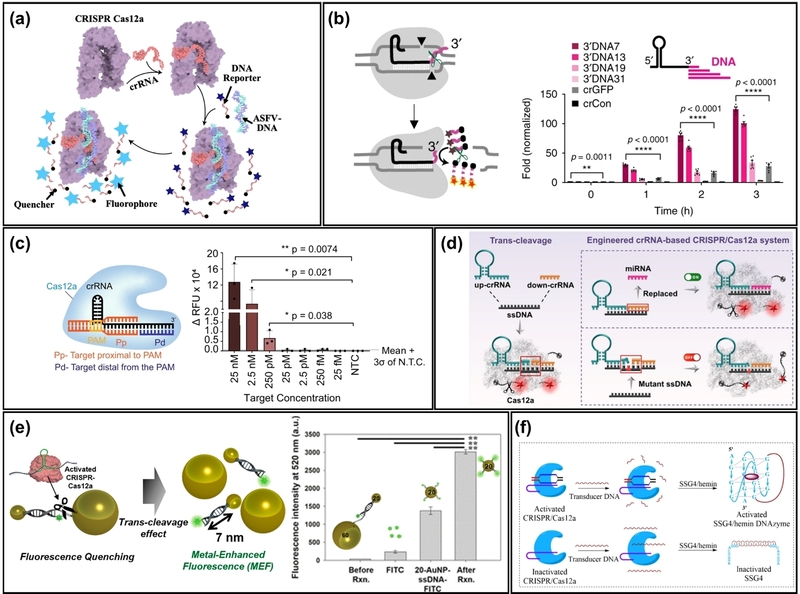
Schematic of Cas12a-based nucleic acid detection. (a) Schematic of a conventional Cas12a-based fluorescence assay for the detection of African swine fever virus (ASFV) DNA utilizing collateral cleavage activity. (b) Engineered crRNA strategy where a 3’-end DNA extension enhances the trans-cleavage activity of Cas12a for improved sensitivity. (c) Split-target recognition strategy allowing Cas12a to detect ssRNA targets by co-localizing two split strands (proximal and distal) on the crRNA. (d) Split-crRNA strategy for miRNA detection, where the target miRNA replaces a segment of the crRNA to activate the Cas12a complex. (e) MEF-based reporter system utilizing trans-cleavage to separate 20 nm and 60 nm AuNPs, thereby restoring and enhancing the fluorescence signal. (f) G-quadruplex-based reporter strategy where Cas12a cleavage regulates the formation of a G-quadruplex/hemin DNAzyme for signal generation. Adapted from (a) Ref. [27], (b) Ref. [29], (c) Ref. [30], (d) Ref. [31], (e) Ref. [32], and (f) Ref. [33].
One effective approach is crRNA engineering. For instance, Nguyen et al. reported that by simply extending the 3’ end of the crRNA with 7-mer DNA, the trans-cleavage activity of the LbCas12a was enhanced 3.5-fold compared to using wild-type crRNA (Fig. 1(b)) [29]. Utilizing this engineered crRNA, they achieved amplification-free detection of the prostate cancer biomarker PCA3 at 25 fM within 6 hours, representing an approximately 400-fold improvement in sensitivity. A similar increase in sensitivity using the engineered crRNA was also observed for the detection of human immunodeficiency virus (HIV) and hepatitis C virus (HCV) genes. crRNA engineering offers various benefits beyond simply increasing sensitivity. Rananaware et al. demonstrated that the trans-cleavage activity of Cas12a is retained when the target strand is split into two short strands that co-localize on the crRNA [30]. The notable point is that the activity of Cas12a is retained even when the short strand at the PAM-distal region is an RNA strand (Fig. 1(c)). This discovery expanded the range of detectable targets to ssRNA, enabling detection down to 132 pM without requiring reverse transcription. Similarly, Zhao et al. confirmed that trans-cleavage activity is maintained when the crRNA’s spacer is split into two short strands that then assemble on a target (Fig. 1(d)) [31]. They also verified that detection is successful when the downstream fragment of the split crRNA (the crRNA at the PAM-distal region) is replaced with a target microRNA (miRNA). Furthermore, both approaches have demonstrated the potential to improve specificity, enabling not only RNA targeting but also effective discrimination of single-base mismatches.
Another effective approach to enhance sensitivity is to utilize a more efficient reporter system than conventional reporter DNA. A Cas12a-based assay typically generates a fluorescent signal by cleaving a fluorophore-quencher dual-tagged DNA. In this system, a fluorophore is initially kept quenched via Förster resonance energy transfer (FRET). Upon cleavage of the DNA, the fluorophore is separated from the quencher, restoring the fluorescent signal. Therefore, enhancing the fluorescent signal change upon cleavage can directly improve sensitivity. In this context, Choi et al. introduced a reporter system integrated with AuNPs to enhance fluorescence signals via metal-enhanced fluorescence (MEF), as shown in Fig. 1(e) [32]. Specifically, they linked two gold nanoparticles (AuNPs) of different sizes (20 nm and 60 nm) with a fluorophore-tagged DNA strand. When a target activates Cas12a, its trans-cleavage activity separates the two nanoparticles, detaching the 60 nm AuNP that quenched the fluorescence. The fluorophore remains at an optimal distance (approx. 7 nm) from the 20 nm AuNP, which in turn enhances the fluorophore’s fluorescence via the MEF effect. Using this reporter system, they successfully detected the breast cancer gene BRCA-1, achieving a detection limit of 0.34 fM within 30 minutes. Similarly, another strategy for improving the reporter system involves the use of G-quadruplexes (Fig. 1(f)) [33,34].
3.2 Cas12a-assisted cascade amplification for nucleic acid detection
CHA is an enzyme-free signal amplification strategy that utilizes the thermodynamically driven hybridization of metastable DNA hairpins. Although CHA offers high specificity and simplicity, its signal amplification is often insufficient to detect ultralow target concentrations. To address this challenge, integrating CHA with the trans-cleavage activity of Cas12a has been suggested as a dual-amplification strategy. Peng et al. designed a target miRNA-initiated CHA circuit to produce dsDNA containing a protospacer sequence, which acts as a dsDNA activator for Cas12a (Fig. 2(a)) [35]. Consequently, a single miRNA target is converted into hundreds of DNA activators, which then trigger Cas12a to cleave thousands of fluorescent reporters per second indiscriminately, achieving two-step signal amplification. This method demonstrated a detection limit of 0.07 fM for miR-21, improving the sensitivity by approximately 6 orders of magnitude compared to conventional CHA assays [14].
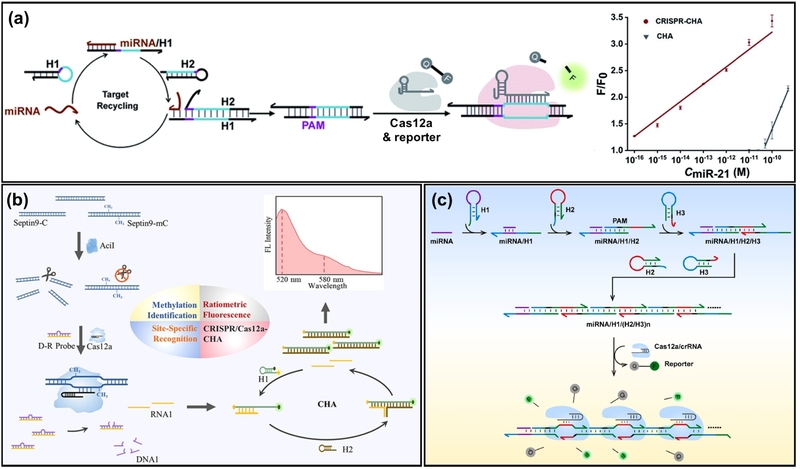
Schematic of Cas12a-assisted cascade amplification for nucleic acid detection. (a) miRNA-initiated CHA strategy where the target triggers a CHA circuit to generate dsDNA products containing PAM sequences, which subsequently activate Cas12a for cascade signal amplification. (b) Cas12a-triggered CHA system for site-specific DNA methylation detection. Activated Cas12a trans-cleaves a DNA-RNA hybrid probe to release an RNA initiator, which triggers a downstream CHA reaction. (c) HCR-integrated Cas12a platform where target miRNA initiates the assembly of long HCR nanowires containing repeating PAM and protospacer units. These nanowires recruit multiple Cas12a complexes, enhancing the cleavage of reporters. Adapted from (a) Ref. [35], (b) Ref. [37], and (c) Ref. [38].
Expanding on this concept, Chen et al. introduced an RNA-based CHA circuit coupled with Cas12a for one-step miRNA detection [36]. This work employs a spacer-blocking strategy in which the crRNA sequence is initially trapped within a hairpin structure. When the target miRNA triggers the CHA circuit, a conformational change in the hairpin exposes the blocked crRNA sequence. Notably, this method exploits the intrinsic cleavage activity of Cas12a to process the pre-crRNA into a mature crRNA, which then guides Cas12a to recognize a DNA activator and cleave reporters. This strategy successfully integrated the preamplification and detection steps into a single-pot reaction, achieving a limit of detection (LOD) of approximately 82 fM.
Conversely, Cas12a can be employed as the initial recognition element for the target, in which its trans-cleavage activity releases a trigger strand to initiate a downstream CHA reaction. This cascade amplification is particularly effective for dsDNA targets. For instance, Ding et al. reported a Cas12a-triggered CHA system for detecting site-specific DNA methylation [37]. In this design, Cas12a is activated by the methylated target DNA and performs trans-cleavage on a DNA-RNA hybrid probe (Fig. 2(b)). The cleavage of the DNA portion releases the short RNA strand, which serves as the initiator for the subsequent CHA cycle. This cascaded amplification achieved a detection limit of 2.02 fM and effectively distinguished colorectal cancer patients from healthy individuals using serum samples.
Similar to CHA, HCR is a prominent enzyme-free signal amplification strategy that utilizes the thermodynamically driven hybridization of metastable DNA hairpins. However, unlike CHA, which operates catalytically to generate multiple discrete DNA duplexes by recycling the initiator strand, HCR is a polymerization process where the initiator strand triggers a cascade of hybridization events, self-assembling into long, nicked double-stranded DNA polymers. This distinct feature of forming high-molecular-weight nanowires enables HCR to concentrate activators spatially or to create physical scaffolds, offering unique advantages when integrated with the CRISPR-Cas12a system. For instance, Jia et al. developed an HCR-integrated Cas12a system for sensitive miRNA detection (Fig. 2(c)) [38]. In this system, a target miRNA initiates the HCR assembly of three hairpins, producing long HCR nanowires that contain repeating units of PAM and protospacer sequences. Consequently, a single miRNA target is converted into a long polymer containing numerous dsDNA activators, which effectively recruit and activate multiple Cas12a/crRNA complexes to cleave reporters, achieving a detection limit of 1 fM for miR-21.
Alternatively, Cas12a can be used to modulate the availability of solid-phase initiators. Ke et al. proposed a chemiluminescence imaging sensor (CLE-CRISPR) where Cas12a activity protects the HCR initiator on magnetic beads [39]. In the absence of the target, the initiator DNA triggers toehold-mediated strand displacement, removing the HCR-initiating sequence (recognition DNA) from the magnetic beads. However, in the presence of the target, activated Cas12a cleaves the initiator DNA, allowing the recognition DNA attached to the beads to initiate HCR. The resulting long DNA polymers bind streptavidin-alkaline phosphatase, resulting in a strong chemiluminescence signal. This strategy achieved an LOD of 3 pM for synthetic DNA.
4. CRISPR-Cas13a
4.1 Cas13a-based nucleic acid detection
Unlike Cas12a, which targets DNA, the CRISPR-Cas13a system specifically recognizes ssRNA. This unique characteristic makes Cas13a a premier candidate for direct RNA detection without the need for reverse transcription. Furthermore, the superior cleavage activity of LbuCas13a has led to recent advances in direct, amplification-free detection. However, relying solely on a single crRNA often limits reaction kinetics, particularly at low target concentrations. To overcome this kinetic bottleneck, Fozouni et al. employed multiple distinct crRNAs to target distinct regions of the viral genome simultaneously (Fig. 3(a)) [25]. This approach effectively increases the concentration of the activated Cas13a-crRNA-target complex, thereby significantly enhancing the fluorescent signal. Leveraging this strategy, they developed an amplification-free SARS-CoV-2 detection method capable of identifying ~100 copies/µL of viral RNA within 30 minutes. Furthermore, by combining this assay with a mobile phone-based fluorescence microscope, they successfully demonstrated a portable readout system suitable for point-of-care testing.
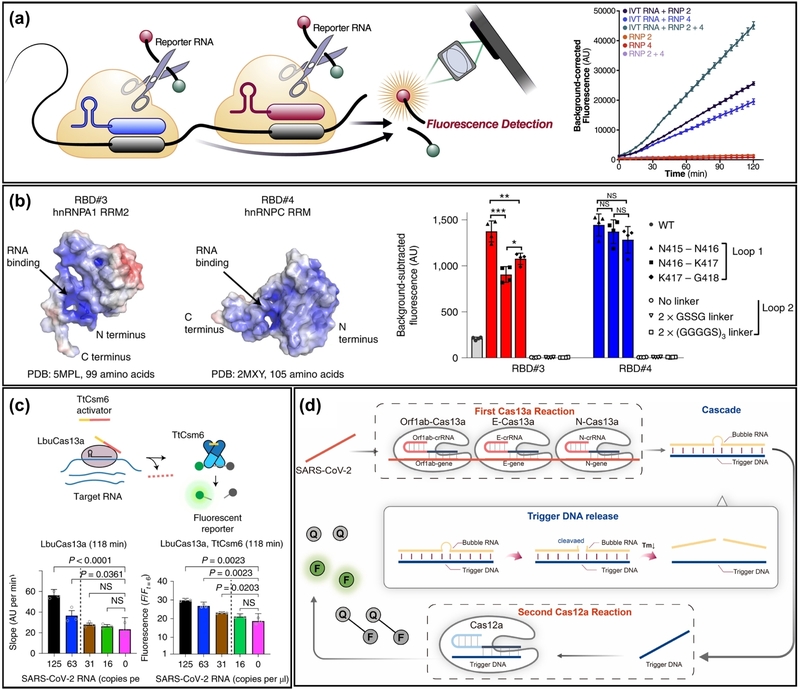
Schematic of Cas13a-based nucleic acid detection and Cas13a-assisted cascade amplification for nucleic acid detection. (a) Multi-crRNA strategy to target different regions of the viral genome simultaneously increases the concentration of activated Cas13a complexes, enhancing the fluorescence signal without amplification. (b) Protein engineering strategy to enhance Cas13a activity. Rational mutagenesis stabilizes the HEPN domain and optimizes RNA binding, resulting in variants (e.g., LwaCas13a-N2V8) with significantly higher turnover rates than the wild type. (c) Cas13a-Csm6 cascade system. Target-activated Cas13a cleaves an RNA linker to release an activator, which subsequently triggers the robust RNase activity of Csm6, achieving sequential signal amplification. (d) ASCas system coupling Cas13a with a CHA circuit. Adapted from (a) Ref. [25], (b) Ref. [40], (c) Ref. [41], and (d) Ref. [42].
Another approach focuses on protein engineering to intrinsically enhance the catalytic efficiency of Cas13a. Yang et al. performed rational mutagenesis on LwaCas13a (Cas13a from Leptotrichia wadei) to improve its collateral cleavage activity [40]. By stabilizing the conformation of the HEPN domain and optimizing RNA-binding interactions, they engineered variants (e.g., LwaCas13a-N2V8) that exhibited significantly higher turnover rates than wild-type LwaCas13a (Fig. 3(b)). This enhanced enzymatic activity enabled ultrasensitive detection of target RNA at low femtomolar levels without nucleic acid amplification, overcoming the inherent sensitivity limitations of native Cas proteins.
4.2 Cas13a-assisted cascade amplification for nucleic acid detection
While optimizing the intrinsic activity of Cas13a significantly improves sensitivity, detecting attomolar concentrations of RNA often requires further signal enhancement. Similar to the strategies employed with Cas12a, the trans-cleavage activity of Cas13a can be utilized as a molecular switch to trigger downstream catalytic or assembly reactions, creating a cascade effect that further amplifies the signal. Liu et al. introduced a cascade system linking Cas13a to Csm6, a Type III CRISPR accessory nuclease (Fig. 3(c)) [41]. In this system, Cas13a recognizes the target RNA and cleaves a specific RNA linker that protects the Csm6 activator. Once the activator is released, it triggers Csm6, which possesses robust RNase activity independent of target binding. This sequential activation—where one target-binding event triggers the activation of multiple turnover enzymes—resulted in an 8-fold improvement in signal kinetics compared to Cas13a alone, enabling detection limits down to the attomolar range.
Alternatively, Cas13a can be integrated with non-enzymatic amplification circuits such as HCR or CHA. Unlike Cas12a, Cas13a cleaves RNA promiscuously; thus, it can be used to remove an RNA lock or cleave a hairpin loop that prevents the initiation of downstream reactions. For instance, Zhang et al. developed an attomolar-level-sensitive cascade system (ASCas) by coupling Cas13a with a CHA circuit (Fig. 3(d)) [42]. In this system, the trans-cleavage activity of Cas13a cleaves the loop region of a specific hairpin probe. This cleavage exposes a previously hidden toehold sequence, which then triggers hybridization with a second hairpin, initiating the CHA cycle. This design demonstrates that the cleavage-mediated activation principle is universally applicable across different Cas effectors to achieve ultrasensitive detection.
5. READOUT STRATEGY
Although fluorescence-based assays, typically using plate readers, are the most common readout methods for CRISPR-Cas diagnostics, recent studies have integrated CRISPR systems with more sensitive readout modalities for detecting low-abundance targets without preamplification.
5.1 Electrochemical assay
Electrochemical biosensors convert biological recognition events into measurable electrical signals (e.g., current, potential, or impedance). This modality is particularly advantageous due to its high sensitivity and rapid response. In a typical CRISPR-based electrochemical assay, single-stranded DNA (for Cas12a) or RNA (for Cas13a) reporters tagged with a redox probe (e.g., methylene blue) are immobilized on an electrode surface. Upon target recognition, the activated Cas nuclease cleaves the reporter, causing the redox probe to diffuse away from the surface and leading to a significant reduction in the electrochemical signal. Ge et al. used Cas12a-mediated cleavage to precisely control the distance between a redox probe and the electrode surface (Fig. 4(a)) [43]. This ratiometric design effectively minimized environmental interference, enabling the detection of genetically modified soybean DNA with an ultralow limit of detection of 0.48 fM within 60 minutes. Sheng et al. introduced an electrochemical assay based on a CHA circuit coupled with Cas13a (Fig. 4(b)) [44]. In this system, the trans-cleavage of Cas13a triggers a downstream CHA, creating a dual-amplification pathway. This strategy achieved an exceptional LOD of 50 aM for tumor-related RNAs within a total assay time of 36 minutes. This result demonstrates that electrochemical readouts can offer both attomolar sensitivity and rapid turnaround times for diverse nucleic acid targets.
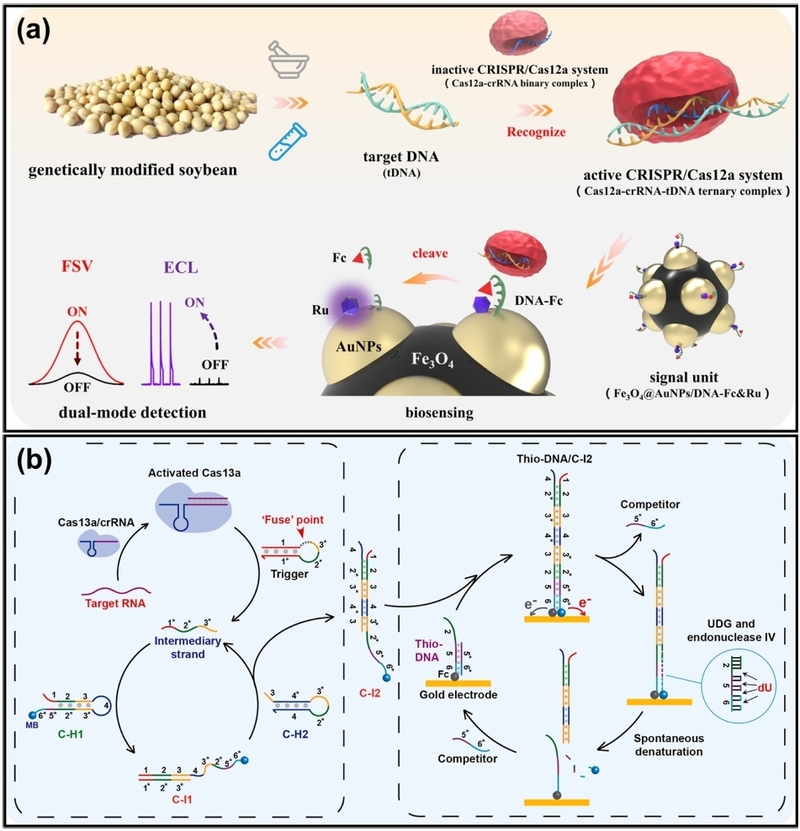
Schematic of electrochemical assay. (a) Cas12a-mediated ratiometric electrochemical sensor for genetically modified soybean detection. The system utilizes Cas12a cleavage to precisely regulate the distance between the redox probe and the electrode surface, minimizing environmental interference for accurate quantification. (b) Cas13a-powered electrochemical assay coupled with a catalytic hairpin assembly (CHA) circuit. Activated Cas13a trans-cleaves a reporter to release a trigger strand, which initiates a downstream CHA reaction on the electrode, creating a dual-amplification pathway for ultrasensitive RNA detection. Adapted from (a) Ref. [43] and (b) Ref. [44].
5.2 Surface-enhanced Raman scattering assay
SERS is a spectroscopic technique that dramatically amplifies the Raman scattering signals of molecules adsorbed on nanostructured metal surfaces via plasmonic enhancement [45-47]. SERS-based readouts offer rich spectral fingerprints, high specificity, and single-molecule sensitivity, making them highly suitable for multiplexing and nucleic acid detection. In a typical CRISPR-SERS assay, Raman dye-labeled reporters are placed near plasmonic surfaces to form hotspots with intense electromagnetic fields. The collateral cleavage activity of Cas proteins detaches the Raman dye from the surface, resulting in a quantifiable change in the SERS signal. For instance, Choi et al. proposed a Cas12a-assisted SERS assay using Raman reporter-functionalized AuNPs on SERS-active graphene oxide (GO)/periodic triangular gold nanoflower (TANF) arrays (Fig. 5(a)) [48]. The TANF arrays and AuNPs serve as hotspots, yielding a significantly enhanced SERS signal when Cas12a is not activated. In the presence of the target DNA, the activation of Cas12a cleaves the ssDNA between GO/TANF and AuNPs, resulting in a decreased Raman signal. This strategy enables an extremely low detection limit of 1 aM without amplification steps, enabling a short assay time of 20 minutes. Similarly, Pang et al. proposed a Cas12a-assisted SERS assay implemented as a lateral flow assay (LFA) (Fig. 5(b)) [49]. They used digoxin antibody-labeled SiO2@Au@5,5′-dithiobis-(2-nitrobenzoic acid) (DTNB) NPs as SERS tags dispersed on the conjugation pad. In the absence of target DNA, Cas12a is inactive, allowing the biotin–DNA–digoxin complex to conjugate with the SERS tag and fix onto the streptavidin-functionalized test line, inducing both strong SERS and colorimetric signals. In the presence of target DNA, activated Cas12a cleaves the biotin–DNA–digoxin complex, preventing the fixation of SERS tags. This assay directly quantifies HIV-1 dsDNA down to 0.3 fM without any preamplification step.
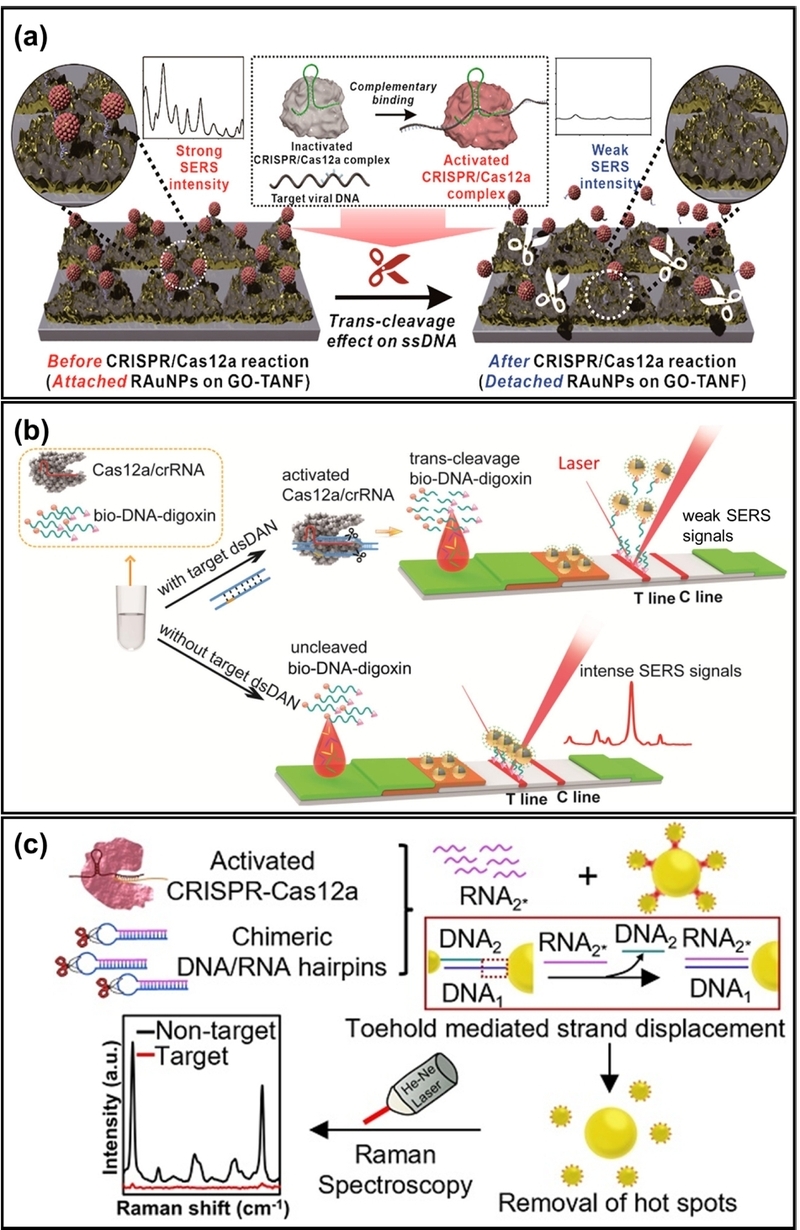
Schematic of electrochemical assay. (a) Cas12a-assisted SERS biosensor utilizing a GO/TANF nanoarray. The trans-cleavage activity of activated Cas12a cleaves the ssDNA linker, detaching Raman reporter-functionalized AuNPs from the surface and reducing the SERS signal. (b) SERS-based lateral flow assay (LFA) for DNA detection. In the presence of the target, activated Cas12a cleaves the biotin–DNA–digoxin complex, preventing the immobilization of SERS tags on the test line and resulting in a signal decrease. (c) Homogeneous colloidal SERS nanoplatform based on plasmonic satellite nanoconjugates. Cas12a cleavage of a chimeric DNA/RNA hairpin releases an RNA strand that displaces satellite AuNPs from the core via toehold-mediated strand displacement, eliminating electromagnetic hotspots. Adapted from (a) Ref. [48], (b) Ref. [49], and (c) Ref. [50].
The realization of CRISPR-SERS assays is not limited to solid substrates; they can also be implemented as homogeneous colloidal assays. Yin et al. proposed a Cas12a-integrated SERS nanoplatform based on plasmonic satellite nanoconjugates (Fig. 5(c)) [50]. To significantly improve sensitivity, they adopted a chimeric DNA/RNA hairpin. The SERS probe consists of a 40 nm AuNP core and Raman reporter-functionalized 13 nm AuNP satellites, linked by partially complementary DNA strands to form hotspots. Upon activation by the target DNA, Cas12a cleaves the loop of the chimeric hairpin, releasing a large amount of RNA. This released RNA is fully complementary to the linker DNA on the core, thereby displacing the satellite nanoparticles. This disassembly destroys the hotspots, leading to a sequential decrease in the SERS signal. This colloidal approach achieved a detection limit down to the attomolar level.
5.3 Digital CRISPR assay
Digital assays represent a paradigm shift from ensemble average measurements to absolute single-molecule quantification. By partitioning the reaction mixture into tens of thousands of picoliter- to femtoliter-sized droplets or microwells using microfluidic technologies, the effective concentration of the target and the reagents is significantly increased within each partition. This confinement effect dramatically enhances reaction kinetics and the signal-to-noise ratio, enabling ultrasensitive detection without the need for preamplification. In amplification-free settings, Tian et al. developed an ultralocalized Cas13a assay that partitions the reaction into picoliter-sized droplets (Fig. 6(a)) [51]. By confining a single target RNA molecule with Cas13a and reporters, they achieved accurate quantification of SARS-CoV-2 RNA and miRNAs with a LOD in the attomolar range, effectively suppressing the background noise observed in bulk reactions. Similarly, Yue et al. applied this droplet microfluidic strategy to the Cas12a system, achieving single-molecule quantification of DNA targets without enzymatic amplification (Fig. 6(b)) [52].
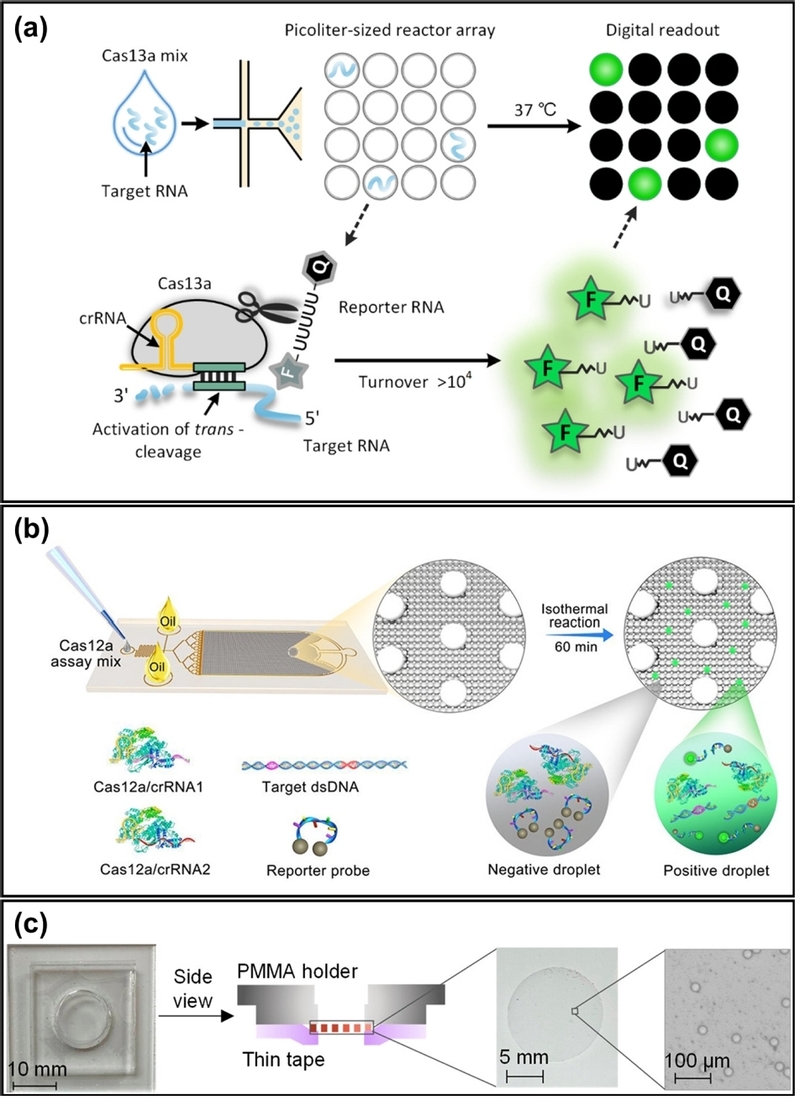
Schematic of digital CRISPR assay. (a) Ultralocalized Cas13a assay using picoliter-sized droplets. Partitioning the reaction into thousands of micro-reactors significantly increases the effective local concentration of targets. (b) Digital Cas12a assay based on droplet microfluidics. This platform partitions the Cas12a reaction mixture into water-in-oil droplets to achieve single-molecule quantification of DNA targets. (c) STAMP platform which utilizes a porous membrane to spontaneously partition the reaction mixture via capillary action. Adapted from (a) Ref. [51], (b) Ref. [52], and (c) Ref. [54].
While droplet microfluidics offers high throughput, it often requires complex pumping systems. To simplify the workflow, physical partitioning methods have been developed. Shinoda et al. introduced SATORI (Single-molecule Assay for Trans-cleavage of Reporter on Integrated chip), which utilizes an array of femtoliter-sized microchambers [53]. This extreme miniaturization further concentrates the fluorescent signal, enabling detection of viral RNA in less than 5 minutes with femtomolar sensitivity. More recently, Nouri et al. proposed the STAMP (Self-digitization through Automated Membrane-based Partitioning) platform (Fig. 6(c)) [54]. This cost-effective method uses a commercially available porous membrane to spontaneously partition the reaction mix via capillary action, eliminating the need for specialized microfluidic chips or pumps while maintaining high sensitivity for HIV-1 viral load monitoring.
6. CONCLUSIONS
This review has highlighted the transformative potential of CRISPR-Cas systems for nucleic acid diagnostics. By circumventing the reliance on complex thermal cycling and expensive enzymatic amplification, CRISPR-based amplification-free strategies offer a paradigm shift toward rapid, simple, and field-deployable testing. As explored in the preceding sections, the trans-cleavage activities of Cas12a and Cas13a serve as the fundamental engine for signal generation, enabling the direct recognition of DNA and RNA targets with high specificity.
A key focus of current research has been bridging the sensitivity gap inherent in amplification-free methods. Integrations of CRISPR systems with non-enzymatic cascade circuits, such as CHA and HCR, have successfully demonstrated attomolar-level sensitivity while maintaining isothermal simplicity. Furthermore, coupling CRISPR systems with advanced readout modalities has significantly improved sensitivity, enabling quantification without preamplification.
Despite these significant advances, several challenges remain for widespread clinical adoption. A primary hurdle is the collateral nature of trans-cleavage, which indiscriminately degrades surrounding reporters. This characteristic poses an inherent challenge for single-pot multiplexing, often requiring spatial separation (e.g., microfluidic chambers) or the use of orthogonal Cas enzymes with distinct cleavage preferences. Additionally, while many assays show promise with synthetic or isolated targets, labor-intensive, time-consuming sample preparation steps remain to extract and purify targets from complex clinical samples. Therefore, their robustness in complex clinical matrices (e.g., whole blood, saliva) requires further validation to prevent false negatives caused by matrix interference or nuclease degradation.
Finally, as these technologies mature from proof-of-concept to standardized commercial platforms, CRISPR-based amplification-free detection is poised to decentralize molecular diagnostics, making precision medicine accessible even in resource-limited settings.
Acknowledgments
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (No. 2022R1C1C2007002).
REFERENCES
-
B.D. Kevadiya, J. Machhi, J. Herskovitz, M.D. Oleynikov, W.R. Blomberg, N. Bajwa, et al., Diagnostics for SARS-CoV-2 infections, Nat. Mater. 20 (2021) 593–605.
[https://doi.org/10.1038/s41563-020-00906-z]

-
A.R. Chandrasekaran, J.A. Punnoose, L. Zhou, P. Dey, B.K. Dey, K. Halvorsen, DNA nanotechnology approaches for microRNA detection and diagnosis, Nucleic Acids. Res. 47 (2019) 10489–10505.
[https://doi.org/10.1093/nar/gkz580]

-
S. Yang, R.E. Rothman, PCR-based diagnostics for infectious diseases: uses, limitations, and future applications in acute-care settings, Lancet Infect. Dis. 4 (2004) 337–348.
[https://doi.org/10.1016/S1473-3099(04)01044-8]

-
W. Liu, L.P. Lee, Toward Rapid and Accurate Molecular Diagnostics at Home, Adv. Mater. 35 (2023) 2206525.
[https://doi.org/10.1002/adma.202206525]

-
W. Liu, F. Yue, L.P. Lee, Integrated Point-of-Care Molecular Diagnostic Devices for Infectious Diseases, Acc. Chem. Res. 54 (2021) 4107–4119.
[https://doi.org/10.1021/acs.accounts.1c00385]

-
J. Cheong, H. Yu, C.Y. Lee, J. Lee, H.-J. Choi, J.-H. Lee, et al., Fast detection of SARS-CoV-2 RNA via the integration of plasmonic thermocycling and fluorescence detection in a portable device, Nat. Biomed. Eng. 4 (2020) 1159–1167.
[https://doi.org/10.1038/s41551-020-00654-0]

-
N.R. Blumenfeld, M.A.E. Bolene, M. Jaspan, A.G. Ayers, S. Zarrandikoetxea, J. Freudman, et al., Multiplexed reverse-transcriptase quantitative polymerase chain reaction using plasmonic nanoparticles for point-of-care COVID-19 diagnosis, Nat. Nanotechnol. 17 (2022) 984–992.
[https://doi.org/10.1038/s41565-022-01175-4]

-
T. AbdElFatah, M. Jalali, S.G. Yedire, I.I. Hosseini, C. del Real Mata, H. Khan, et al., Nanoplasmonic amplification in microfluidics enables accelerated colorimetric quantification of nucleic acid biomarkers from pathogens, Nat. Nanotechnol. 18 (2023) 922–932.
[https://doi.org/10.1038/s41565-023-01384-5]

-
N. Panpradist, E.C. Kline, R.G. Atkinson, M. Roller, Q. Wang, I.T. Hull, et al., Harmony COVID-19: A ready-to-use kit, low-cost detector, and smartphone app for point-of-care SARS-CoV-2 RNA detection, Sci. Adv. 7 (2024) eabj1281.
[https://doi.org/10.1126/sciadv.abj1281]

-
G. Xun, S.T. Lane, V.A. Petrov, B.E. Pepa, H. Zhao, A rapid, accurate, scalable, and portable testing system for COVID-19 diagnosis, Nat. Commun. 12 (2021) 2905.
[https://doi.org/10.1038/s41467-021-23185-x]

-
D. Kang, S. Choi, S. Roh, T. Lee, G. Lee, Recent Advances in Loop-Mediated Isothermal Amplification (LAMP) for On-Site Virus Detection, J. Sens. Sci. Technol. 34 (2025) 432–444.
[https://doi.org/10.46670/JSST.2025.34.5.432]

-
J. Shin, S. Lim, T. Yoon, J. You, Self-Assembly of Rolling Circle Amplification DNA into Nano-particles using Mg2+ and ppi, J. Sens. Sci. Technol. 34 (2025) 399–409.
[https://doi.org/10.46670/JSST.2025.34.5.399]

-
Z. Ge, M. Lin, P. Wang, H. Pei, J. Yan, J. Shi, et al., Hybridization Chain Reaction Amplification of MicroRNA Detection with a Tetrahedral DNA Nanostructure-Based Electrochemical Biosensor, Anal. Chem. 86 (2014) 2124–2130.
[https://doi.org/10.1021/ac4037262]

-
J.C. Park, H. Na, S. Choi, H. Jeon, Y.S. Nam, Target-Catalyzed Self-Assembly of DNA-Streptavidin Nanogel for Enzyme-Free miRNA Assay, Adv. Healthc. Mater. 12 (2023) 2202076.
[https://doi.org/10.1002/adhm.202202076]

-
C.A. Hong, J.C. Park, H. Na, H. Jeon, Y.S. Nam, Short DNA-catalyzed formation of quantum dot-DNA hydrogel for enzyme-free femtomolar specific DNA assay, Biosens. Bioelectron. 182 (2021) 113110.
[https://doi.org/10.1016/j.bios.2021.113110]

-
H. Na, B.I. Koo, J.C. Park, J. Lim, Y. Kim, H.J. Chung, et al., Live-Cell Imaging of MicroRNA Expression via Photoinduced Electron Transfer Controlled by Catalytic Hairpin Assembly, Adv. Healthc. Mater. 13 (2024) 2401483.
[https://doi.org/10.1002/adhm.202401483]

-
B. Li, A.D. Ellington, X. Chen, Rational, modular adaptation of enzyme-free DNA circuits to multiple detection methods, Nucleic Acids Res. 39 (2011) e110.
[https://doi.org/10.1093/nar/gkr504]

-
A. East-Seletsky, M.R. O’Connell, S.C. Knight, D. Burstein, J.H.D. Cate, R. Tjian, et al., Two distinct RNase activities of CRISPR-C2c2 enable guide-RNA processing and RNA detection, Nature 538 (2016) 270–273.
[https://doi.org/10.1038/nature19802]

-
J.S. Gootenberg, O.O. Abudayyeh, J.W. Lee, P. Essletzbichler, A.J. Dy, J. Joung, et al., Nucleic acid detection with CRISPR-Cas13a/C2c2, Science 356 (2017) 438–442.
[https://doi.org/10.1126/science.aam9321]

-
H. Nishimasu, F.A. Ran, P.D. Hsu, S. Konermann, S.I. Shehata, N. Dohmae, et al., Crystal Structure of Cas9 in Complex with Guide RNA and Target DNA, Cell 156 (2014) 935–949.
[https://doi.org/10.1016/j.cell.2014.02.001]

-
C. Anders, O. Niewoehner, A. Duerst, M. Jinek, Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease, Nature 513 (2014) 569–573.
[https://doi.org/10.1038/nature13579]

-
R. Hajian, S. Balderston, T. Tran, T. deBoer, J. Etienne, M. Sandhu, et al., Detection of unamplified target genes via CRISPR–Cas9 immobilized on a graphene field-effect transistor, Nat. Biomed. Eng. 3 (2019) 427–437.
[https://doi.org/10.1038/s41551-019-0371-x]

-
J.S. Chen, E. Ma, L.B. Harrington, M. Da Costa, X. Tian, J.M. Palefsky, et al., CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity, Science 360 (2018) 436–439.
[https://doi.org/10.1126/science.aar6245]

-
S.-Y. Li, Q.-X. Cheng, J.-K. Liu, X.-Q. Nie, G.-P. Zhao, J. Wang, CRISPR-Cas12a has both cis- and trans-cleavage activities on single-stranded DNA, Cell Res. 28 (2018) 491–493.
[https://doi.org/10.1038/s41422-018-0022-x]

-
P. Fozouni, S. Son, M. Díaz de León Derby, G.J. Knott, C.N. Gray, M.V. D’Ambrosio, et al., Amplification-free detection of SARS-CoV-2 with CRISPR-Cas13a and mobile phone microscopy, Cell 184 (2021) 323–333.
[https://doi.org/10.1016/j.cell.2020.12.001]

-
R. Nouri, M. Dong, A.J. Politza, W. Guan, Figure of Merit for CRISPR-Based Nucleic Acid-Sensing Systems: Improvement Strategies and Performance Comparison, ACS Sens. 7 (2022) 900–911.
[https://doi.org/10.1021/acssensors.2c00024]

-
Q. He, D. Yu, M. Bao, G. Korensky, J. Chen, M. Shin, et al., High-throughput and all-solution phase African Swine Fever Virus (ASFV) detection using CRISPR-Cas12a and fluorescence based point-of-care system, Biosens. Bioelectron. 154 (2020) 112068.
[https://doi.org/10.1016/j.bios.2020.112068]

-
S.-Y. Li, Q.-X. Cheng, J.-M. Wang, X.-Y. Li, Z.-L. Zhang, S. Gao, et al., CRISPR-Cas12a-assisted nucleic acid detection, Cell Discov. 4 (2018) 20.
[https://doi.org/10.1038/s41421-018-0028-z]

-
L.T. Nguyen, B.M. Smith, P.K. Jain, Enhancement of trans-cleavage activity of Cas12a with engineered crRNA enables amplified nucleic acid detection, Nat. Commun. 11 (2020) 4906.
[https://doi.org/10.1038/s41467-020-18615-1]

-
S.R. Rananaware, E.K. Vesco, G.M. Shoemaker, S.S. Anekar, L.S.W. Sandoval, K.S. Meister, et al., Programmable RNA detection with CRISPR-Cas12a, Nat. Commun. 14 (2023) 5409.
[https://doi.org/10.1038/s41467-023-41006-1]

-
S. Zhao, Q. Zhang, J. Sun, S. Li, S. Wang, D. Zhou, et al., Simply Engineered crRNA with CRISPR/Cas12a System Enables Wide-Scope Nucleic Acid Biomarker Analysis, Nano Lett. 25 (2025) 8431–8441.
[https://doi.org/10.1021/acs.nanolett.5c01939]

-
J.H. Choi, J. Lim, M. Shin, S.H. Paek, J.W. Choi, CRISPR-Cas12a-Based Nucleic Acid Amplification-Free DNA Biosensor via Au Nanoparticle-Assisted Metal-Enhanced Fluorescence and Colorimetric Analysis, Nano Lett. 21 (2021) 693–699.
[https://doi.org/10.1021/acs.nanolett.0c04303]

-
Y. Chen, B. Lv, W. Wang, Y. Wu, D. Li, Structure-switching G-quadruplex: An efficient CRISPR/Cas12a signal reporter for label-free colorimetric biosensing, Int. J. Biol. Macromol. 307 (2025) 142410.
[https://doi.org/10.1016/j.ijbiomac.2025.142410]

-
X. Zhao, Y. Zhou, X. Lv, H. Zhang, X. Cui, T. Gong, et al., G-quadruplex molecular beacon: A versatile CRISPR/Cas12a reporter for rapid and label-free biosensing, Sens. Actuators B Chem. 380 (2023) 133310.
[https://doi.org/10.1016/j.snb.2023.133310]

-
S. Peng, Z. Tan, S. Chen, C. Lei, Z. Nie, Integrating CRISPR-Cas12a with a DNA circuit as a generic sensing platform for amplified detection of microRNA, Chem. Sci. 11 (2020) 7362–7368.
[https://doi.org/10.1039/D0SC03084H]

-
P. Chen, L. Wang, P. Qin, B.C. Yin, B.C. Ye, An RNA-based catalytic hairpin assembly circuit coupled with CRISPR-Cas12a for one-step detection of microRNAs, Biosens. Bioelectron. 207 (2022) 114152.
[https://doi.org/10.1016/j.bios.2022.114152]

-
L. Ding, S. Cao, C. Qu, Y. Wu, S. Yu, Ratiometric CRISPR/Cas12a-Triggered CHA System Coupling with the MSRE to Detect Site-Specific DNA Methylation, ACS Sens. 9 (2024) 1877–1885.
[https://doi.org/10.1021/acssensors.3c02571]

-
H.Y. Jia, H.L. Zhao, T. Wang, P.R. Chen, B.C. Yin, B.C. Ye, A programmable and sensitive CRISPR/Cas12a-based MicroRNA detection platform combined with hybridization chain reaction, Biosens. Bioelectron. 211 (2022) 114382.
[https://doi.org/10.1016/j.bios.2022.114382]

-
X. Ke, Y. Ou, Y. Lin, T. Hu, Enhanced chemiluminescence imaging sensor for ultrasensitive detection of nucleic acids based on HCR-CRISPR/Cas12a, Biosens. Bioelectron. 212 (2022) 114428.
[https://doi.org/10.1016/j.bios.2022.114428]

-
J. Yang, Y. Song, X. Deng, J.A. Vanegas, Z. You, Y. Zhang, et al., Engineered LwaCas13a with enhanced collateral activity for nucleic acid detection, Nat. Chem. Biol. 19 (2023) 45–54.
[https://doi.org/10.1038/s41589-022-01135-y]

-
T.Y. Liu, G.J. Knott, D.C.J. Smock, J.J. Desmarais, S. Son, A. Bhuiya, et al., Accelerated RNA detection using tandem CRISPR nucleases, Nat. Chem. Biol. 17 (2021) 982–988.
[https://doi.org/10.1038/s41589-021-00842-2]

-
Y. Zhang, Y. Chen, Q. Zhang, Y. Liu, X. Zhang, An aM-level sensitive cascade CRISPR-Dx system (ASCas) for rapid detection of RNA without pre-amplification, Biosens. Bioelectron. 230 (2023) 115248.
[https://doi.org/10.1016/j.bios.2023.115248]

-
H. Ge, X. Wang, J. Xu, H. Lin, H. Zhou, T. Hao, et al., A CRISPR/Cas12a-Mediated Dual-Mode Electrochemical Biosensor for Polymerase Chain Reaction-Free Detection of Genetically Modified Soybean, Anal. Chem. 93 (2021) 14885–14891.
[https://doi.org/10.1021/acs.analchem.1c04022]

-
Y. Sheng, T. Zhang, S. Zhang, M. Johnston, X. Zheng, Y. Shan, et al., A CRISPR/Cas13a-powered catalytic electrochemical biosensor for successive and highly sensitive RNA diagnostics, Biosens. Bioelectron. 178 (2021) 113027.
[https://doi.org/10.1016/j.bios.2021.113027]

-
R. Park, S. Lee, H.S. Jung, Direct Surface Growth of Plasmonic Nanomaterials: A Novel Approach for SERS Sensing Applications, J. Sens. Sci. Technol. 34 (2025) 445–457.
[https://doi.org/10.46670/JSST.2025.34.5.445]

-
M. Ku, S. Lee, Y.W. Han, Advanced Strategies for SERS Design as a Next-Generation Biosensing Platform: A Mini Review, J. Sens. Sci. Technol. 34 (2025) 314–323.
[https://doi.org/10.46670/JSST.2025.34.4.314]

-
K. Chai, H. Park, J. Park, Fabrication of Nanopillars-Based Raman Substrates for Maximizing SERS Effects: Focusing on Advancement and Applications, J. Sens. Sci. Technol. 34 (2025) 67–75.
[https://doi.org/10.46670/JSST.2025.34.2.67]

-
J.H. Choi, M. Shin, L. Yang, B. Conley, J. Yoon, S.N. Lee, et al., Clustered Regularly Interspaced Short Palindromic Repeats-Mediated Amplification-Free Detection of Viral DNAs Using Surface-Enhanced Raman Spectroscopy-Active Nanoarray, ACS Nano 15 (2021) 13475–13485.
[https://doi.org/10.1021/acsnano.1c03975]

-
Y. Pang, Q. Li, C. Wang, S. Zhen, Z. Sun, R. Xiao, CRISPR-cas12a mediated SERS lateral flow assay for amplification-free detection of double-stranded DNA and single-base mutation, Chem. Eng. J. 429 (2022) 132109.
[https://doi.org/10.1016/j.cej.2021.132109]

-
B. Yin, Q. Zhang, X. Xia, C. Li, W.K.H. Ho, J. Yan, et al., A CRISPR-Cas12a integrated SERS nanoplatform with chimeric DNA/RNA hairpin guide for ultrasensitive nucleic acid detection, Theranostics 12 (2022) 5914–5930.
[https://doi.org/10.7150/thno.75816]

-
T. Tian, B. Shu, Y. Jiang, My Ye, L. Liu, Z. Guo, et al., An Ultralocalized Cas13a Assay Enables Universal and Nucleic Acid Amplification-Free Single-Molecule RNA Diagnostics, ACS Nano 15 (2021) 1167–1178.
[https://doi.org/10.1021/acsnano.0c08165]

-
H. Yue, B. Shu, T. Tian, E. Xiong, M. Huang, D. Zhu, et al., Droplet Cas12a Assay Enables DNA Quantification from Unamplified Samples at the Single-Molecule Level, Nano Lett. 21 (2021) 4643–4653.
[https://doi.org/10.1021/acs.nanolett.1c00715]

-
H. Shinoda, Y. Taguchi, R. Nakagawa, A. Makino, S. Okazaki, M. Nakano, et al., Amplification-free RNA detection with CRISPR–Cas13, Commun. Biol. 4 (2021) 476.
[https://doi.org/10.1038/s42003-021-02001-8]

-
R . Nouri, Y. Jiang, A.J. Politza, T. Liu, W.H. Greene, Y. Zhu, et al., STAMP-Based Digital CRISPR-Cas13a for Amplification-Free Quantification of HIV-1 Plasma Viral Loads, ACS Nano 17 (2023) 10701–10712.
[https://doi.org/10.1021/acsnano.3c01917]


Jae Chul Park is a Postdoctoral fellow in Center for Advanced Biomolecular Recognition at Korea Institute of Science and Technology (KIST). He received his B.S., M.S., and Ph.D. degree in materials science and engineering from Korea Advanced Institute of Science and Technology (KAIST) in 2013, 2015, and 2021. He then moved to KIST as a postdoctoral fellow in Center for Advanced Biomolecular Recognition. His research interests include synthesis, processing, and self-assembly of nanomaterials for biosensing application.
