
Advanced Strategies for SERS Design as a Next-Generation Biosensing Platform: A Mini Review
ⓒ The Korean Sensors Society
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License(https://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
As human lifespans continue to increase, the demand for faster and more accurate medical diagnostics is also rising. Biosensors have evolved accordingly, and point-of-care systems such as rapid antigen tests and glucose monitors are now widely adopted. However, the range of detectable biomolecules remains limited due to inadequate sensitivity and specificity. Among emerging technologies, surface-enhanced Raman spectroscopy (SERS) shows strong potential because of its exceptional sensitivity, specificity, and speed. SERS chips enable direct detection without sample pretreatment, offering significant advantages over conventional methods. Nonetheless, challenges remain in quantification and in distinguishing between structurally similar targets. To address these limitations, recent research has focused on structural innovations—particularly nano-patterning and surface functionalization. These strategies improve reproducibility and analytical precision, positioning SERS as a promising platform for next-generation biosensors. With continued advancement, SERS-based systems are poised to shape the future of medical diagnostics and play a key role in the Fourth Industrial Revolution.
Keywords:
Surface-enhanced Raman spectroscopy (SERS), Internal standard, Electric field extension, Surface functionalization1. INTRODUCTION
Raman scattering arises from characteristic molecular vibrations induced by inelastic photon scattering. As a result, Raman spectroscopy (RS) has become a widely used analytical technique due to its rapid, non-invasive, and highly specific detection capabilities [1,2]. However, its practical application in molecular sensing has been historically limited by its inherently weak signal. RS exhibits a low scattering cross-section, approximately 10-31 cm² sr-1, making it difficult to detect small quantities of analytes [1].
The surface-enhanced Raman spectroscopy (SERS) effect was first observed in the 1970s by Fleischmann et al. [3], marking a significant advancement in the field of RS. This phenomenon involved a notable enhancement of Raman scattering signals when molecules were positioned on a silver electrode. Subsequent research by Jeanmaire and Van Duyne [4] further elucidated the underlying electromagnetic mechanisms contributing to this enhancement.
Since the discovery of SERS, it has emerged as a promising candidate for next-generation sensor platforms, particularly in biosensing applications [5-7]. Advances in nanotechnology have enabled the fabrication of nanostructures capable of generating enhanced electric fields under laser excitation [2,8,9]. The region exhibiting the greatest electric field enhancement, known as the "hot spot," largely determines the strength of the SERS effect [8,9]. With the development of precise hot spot engineering techniques, even single-molecule detection has become possible [10]. This high sensitivity is especially beneficial in biosensing, enabling the accurate detection of trace biomarkers for early disease diagnosis.
However, to advance SERS biosensors from being merely promising to truly practical, several challenges must be addressed. The first involves hot spots, which often exhibit inconsistent sizes and distributions. This inconsistency leads to fluctuations in Raman intensities across a single substrate, resulting in non-uniform measurements and reduced accuracy in quantitative analysis. Second, due to their large size, biomolecules often cannot fully access the hot spots, limiting their exposure to the SERS effect and thereby reducing overall sensitivity. Third, target bio-analytes typically coexist with various non-target biomolecules, making it difficult to distinguish target analytes on the substrate, as signal interference from other substances may occur.
This study reviews approaches to overcoming these challenges, with a primary focus on advanced strategies for innovating the structural design of SERS substrates. First, we provide an overview of the basic principles of SERS and its function as a biosensor. Next, we examine current SERS nanostructure designs that incorporate internal standard materials, extend the range of electric fields, and functionalize surfaces for selective adsorption of target analytes. Finally, we offer perspectives on enhancing its performance as an effective biosensor.
2. FUNDAMENTALS OF SERS-BASED BIOSENSORS
2.1 Principle of the SERS
Two primary mechanisms have been proposed to explain the amplification of Raman signals: the electromagnetic (EM) and chemical enhancement (CE) mechanisms. In the EM mechanism, Raman signal enhancement results from the electric field generated by localized surface plasmon resonance (LSPR) on SERS substrates made of noble metals such as gold and silver (Fig. 1 (a)). When light of a specific wavelength is incident, the free electrons on the metal surface oscillate—a phenomenon known as LSPR—producing a strong electric field [11,12]. If the gap between the metal structures is extremely narrow, typically between 1 and 5 nm, the electric field intensifies further, significantly increasing Raman intensity [8]. The region where this concentrated electric field forms is known as the hot spot [13].
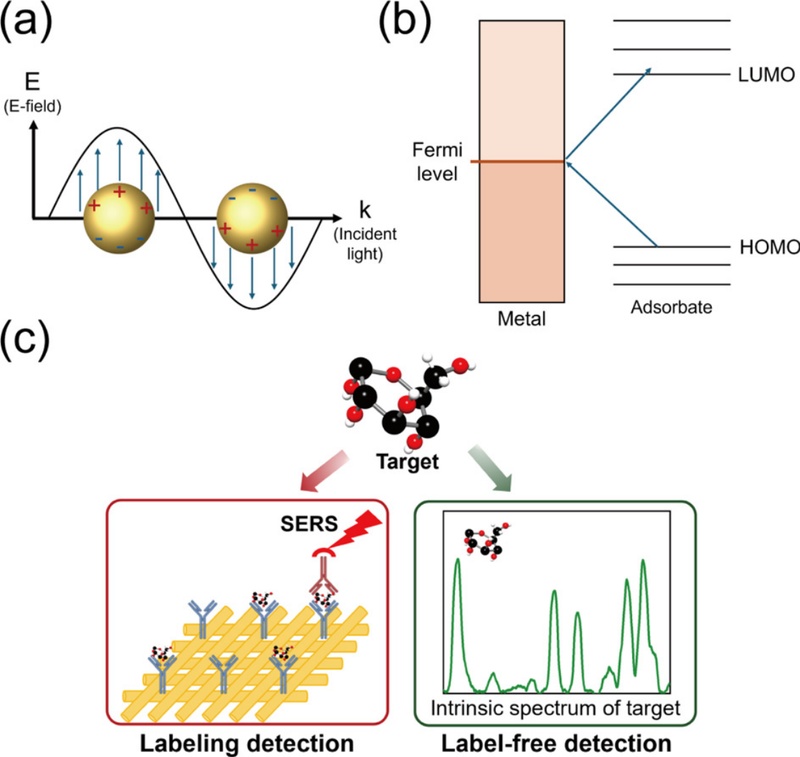
Principle of the SERS and biosensing strategies. (a) Electromagnetic mechanism and (b) chemical enhancement mechanism of the SERS. (c) Schematic of labeling and label-free approaches for biomolecule detection using SERS.
In contrast, the CE mechanism involves chemical interaction between the analyte and SERS substrate (Fig. 1 (b)). When analyte molecules are adsorbed onto the SERS structure, charge transfer (CT) can occur between the Fermi level of the metal and the highest occupied molecular orbital (HOMO) and lowest unoccupied molecular orbital (LUMO) of the analyte [14]. If the laser wavelength matches the electronic transition energy of the analyte–SERS complex, resonance Raman scattering is induced, leading to signal amplification [15]. However, CE typically yields a lower enhancement factor of approximately103 compared to the ~105 to 109-fold enhancement achieved by the EM mechanism [8].
Considering these mechanisms, the degree of Raman signal enhancement is closely linked to the shape, composition, and electron density of the SERS-active metal nanostructures [16]. Therefore, designing nanostructures with densely packed and highly efficient hot spots is essential for detecting trace levels of biomolecules using SERS.
2.2 Detection Strategies of the SERS-based Biosensor
Two main strategies are commonly used for SERS-based biosensing: labeling and label-free methods (Fig. 1 (c)). The labeling approach employs a Raman reporter that generates strong Raman signals, enabling indirect detection of the target biomolecule [17]. This method is applied when the target molecule has a low Raman cross-section or cannot be efficiently positioned at the hot spot. Effective reporters should have a high Raman cross-section, a simple structure with few Raman peaks, resistance to photobleaching, and strong affinity for the target molecule [18].
The labeling method, however, presents challenges in quantifying target analytes and involves several drawbacks, including longer processing times, larger reaction volumes, and the need for additional steps [19]. In contrast, the label-free method directly monitors the Raman spectra of analytes through their interaction with the SERS structure [20]. Nevertheless, detecting and quantifying trace amounts of biomolecules remains difficult due to their inherently low Raman cross-section and potential interference from other metabolites [21]. Therefore, to fully leverage the versatility of label-free methods across various biomolecules, additional strategies are required to enhance biosensing performance—particularly in specificity, sensitivity, and quantification accuracy.
3. STRATEGIES FOR DESIGN OF SERS BIOSENSORS
3.1 Incorporation of Internal Standard
Owing to variations in hot spot distribution and fluctuations in instrument conditions, Raman intensity can differ significantly across different locations on a single substrate. This presents a major challenge for the accurate quantification of specific analyte concentrations. Recently, researchers have incorporated internal standard (IS) materials into SERS substrates and used the Raman intensity ratio (analyte intensity (Ia) / IS intensity (IIS)) to quantify the analyte [22]. The IS is incorporated to ensure that both the Raman signals of the internal standard and target analyte are simultaneously affected by the same instrumental conditions and hot spots at a measurement location. Therefore, despite significant fluctuations in Ia, the normalized Ia/IIS ratio can remain consistent, as IIS fluctuates in the same manner as Ia.
A particular advantage of incorporating IS is the extensive diversity in both material and structure selection. Theoretically, it is possible to employ any material with a distinct Raman peak that does not overlap with the characteristic peaks of the target analyte. Various materials including inorganic dyes, silicon, graphene, polydimethylsiloxane (PDMS), and even biological entities such as bacteria have been effectively utilized as ISs [23-33]. These materials are designed in structures that allow them to be influenced by the hot spots, while avoiding competition with the analytes for adsorption and Raman signal emission.
One of the most widely investigated approaches is to ‘embed’ the IS material in the metal nanostructures, while isolating it from the analytes. Li et al. [23] designed an array of Janus gold particles, embedding 4-mercaptopyridine (4-Mpy) as the IS on one side of each particle (Fig. 2 (a)). The 4-Mpy was simultaneously separated from the target analyte, Rhodamine 6G (R6G), which was adsorbed on the opposite side. The relative standard deviation (RSD) of the intensity ratio was 5.3%, showing a significant improvement in spot-to-spot variation with IS calibration compared to the 16% RSD observed for intensity values without calibration. Additionally, the benefit of separating R6G from 4-Mpy was confirmed using a quantification plot, which showed an R2 of 0.995 when 4-Mpy was measured separately from R6G, compared to an R2 of 0.988 when the two were mixed (Fig. 2 (b)).
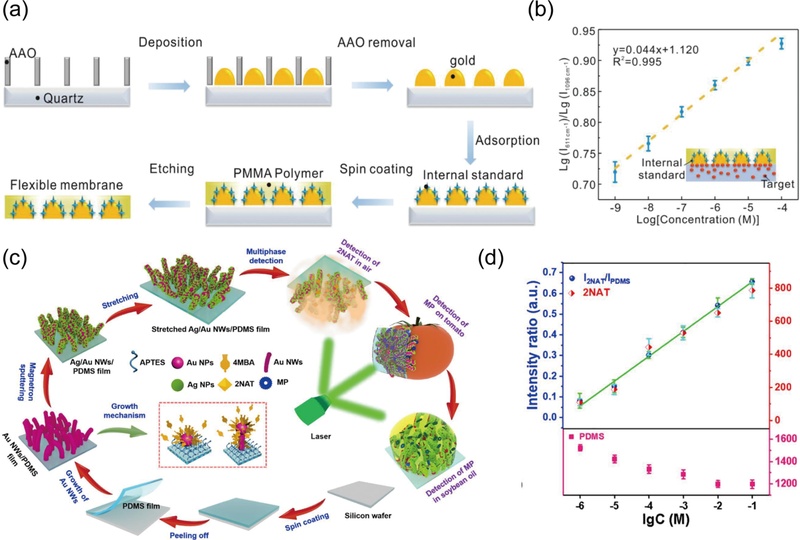
Internal Standard (IS) Incorporation. (a) IS (4-Mpy) embedding and isolating from the analyte (R6G) through the fabrication of Janus gold nanoparticles. (b) Quantification of R6G based on the intensity ratio of IR6G/I4-Mpy. Reproduced with permission from ref. [23]. Copyright (2023) Wiley-VCH GmbH. (c) PDMS film as the intrinsic IS and Ag/Au nanowire growth substrate for Raman measurements on 2-NAT and MP molecules. (d) Quantification of 2-NAT using the intensity ratio of I2-NAT/IPDMS (blue) and plotting of I2-NAT (red) and IPDMS (pink) values against 2-NAT concentration. Reproduced with permission from ref [31]. Copyright (2020) Elsevier.
Nevertheless, the possibility of competition between analytes and the IS, the complexity of structural design, and the natural aggregation of particles pose challenges to the sensitivity and reproducibility of embedded-IS fabrications [28-30]. To address these challenges, the ‘intrinsic IS’ method has been introduced, where the substrate material beneath the metal nanostructure serves as the IS, thus eliminating the need for additional molecules for calibration. Ma et al. [31] utilized a PDMS film not only as a substrate for Ag/Au nanowire growth, but also as the IS, demonstrating quantification of methyl parathion (MP) on food and 2-naphthylamine gas (2-NAT) (Fig. 2 (c)). By comparing the quantification results using the Raman intensities (I2-NAT) and intensity ratios (I2-NAT/IPDMS), the improvement in plot linearity with IS calibration was verified with R2 values of 0.976 and 0.998, respectively (Fig. 2 (d)). In particular, the consistent decrease in IPDMS with increasing 2-NAT concentration showed its effectiveness in calibration.
This highlighted intrinsic IS as a seamlessly integrated calibrator, enhancing reliability without complicated fabrication steps.
3.2 Extension of Electric Field Area
One of the most critical limitations of SERS biosensing is the extremely small hot spots. It is almost impossible to detect biomolecules such as proteins that are bigger than hot spots. Recently, researchers have used nano-optics on SERS such as waveguide-enhanced Raman spectroscopy (WERS), photonic crystal (PhC) SERS, and other techniques. By utilizing nanostructures based on nano-optics, it is possible to overcome the limitations of small hot spot sizes through electric field propagation or electric field expansion.
Recently, Fu et al. [34] demonstrated Waveguide-SERS (W-SERS), which was used to observe plasmonic mode propagation along the nanostructure [34]. In another study about waveguide-enhanced SERS, Dhakal et al. [35] demonstrated nanophotonic waveguide-enhanced Raman spectroscopy (NWERS). NWERS relies on enhancing the excitation and collection of spontaneous Raman scattering near nanophotonic waveguides. As shown in Fig. 3 (a), the waveguide system was composed of Si3N4 circuits, with the top inset showing a generic slot waveguide. The refractive index values were n1=1.45 (SiO2), n2=1.89 (Si3N4), n3=1.33 (H2O), and h=220 nm. The waveguide nanostructure was compared between strip type and slot type through the power map calculated by the COMSOL finite elements mode solver for the TE mode (Figs. 3 (b) and 3 (c)). The total width of the slot waveguide was 850 nm with a 150 nm gap and the strip waveguide was 600 nm wide without a gap. Through the power map, the maximum power area was extended in the gap of the slot waveguide (Fig. 3 (c)). Thus, the slot waveguide in the integrated system was utilized and the measurement ability was compared with the commercial Raman confocal microscope (Fig. 3 (d)). The spectra of a Rhodamine mono-layer was measured and proved that Raman enhancement was significantly improved with the waveguide structure despite the less power and shorter detection time. Consequently, it was shown that a Si3N4 waveguide is expected to yield a signal 4–5 orders of magnitude higher than that in free space. Additionally, improvements can be achieved by utilizing waveguides with a higher index contrast and by optimizing waveguide designs for enhanced efficiency.
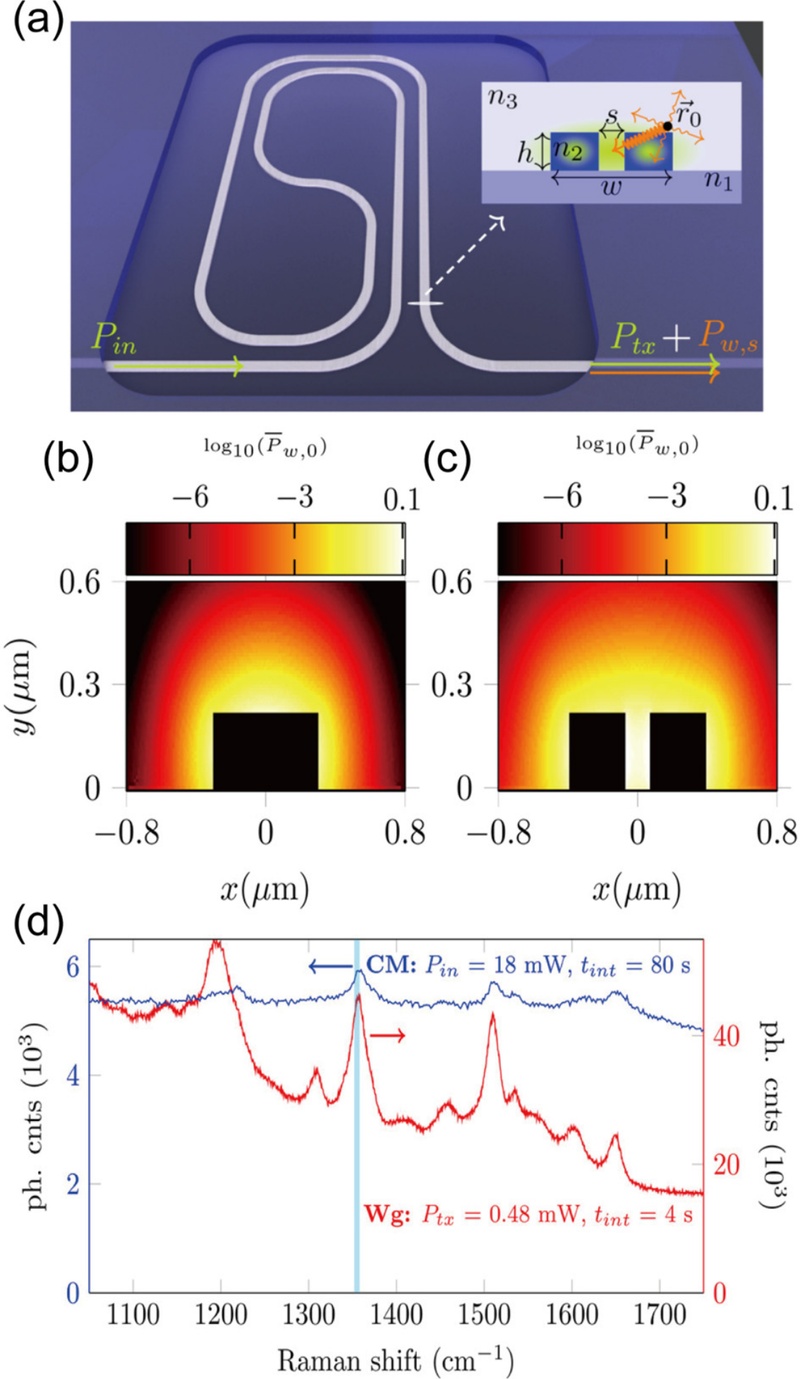
Waveguide-enhanced SERS. (a) Schematic of the waveguide system. Normalized collected power for (b) strip-type and (c) slot-type waveguides. (d) Raman spectra of Rhodamine measured using a commercial Raman microscope (CM) and a slot waveguide (Wg). Reproduced with permission from ref. [35]. Copyright (2016) American Chemical Society.
Another strategy for the expansion of hot spots is to utilize the PhC structure. The key features of PhC are the sub-wavelength scaled periodicity for the periodic potential and refractive index. Gaponenki et al. [36] defined 1 to 3-dimensional periodic PhC structures such as a comb-like configuration (1-D), hexagonal-like configuration (2-D), and woodpile-like configuration (3-D). Accordingly, the PhC-SERS structure demonstrated by Kim et al. [37] was classified as a 2-D structure [37]. As illustrated in Fig. 4 (a), they designed the PhC-SERS structure with Ag nanoparticles on SiO2 nanorods including the TiO2, SiO2, and epoxy grating layer below. The scanning electron microscopy (SEM) image in Fig. 4 (b) shows randomly distributed nanorods with a height of 50 nm. The cross-sectional electric field distribution was compared between the structure of Ag nanoparticles on the glass only (GL-SERS, Fig. 4 (c)) and on the PhC structure depicted in Fig. 4 (a) (PhC-SERS, Fig. 4 (d)). It was verified that the extended electric field area occurred only in PhC-SERS. Furthermore, the Raman enhancement of each structure was investigated to prove the SERS effect of the PhC structure by measuring 1,2-bis(4-pyridyl)ethylene (BPE) (Fig. 4 (e)). In consistency with the simulated results, the Raman enhancement of PhC-SERS with resonance was the highest among GL-SERS and PhC-SERS with and without resonance. The inset of Fig. 4 (e) shows the SERS effect of PhC-SERS enabling the detection of less than 1 femto mole of BPE.
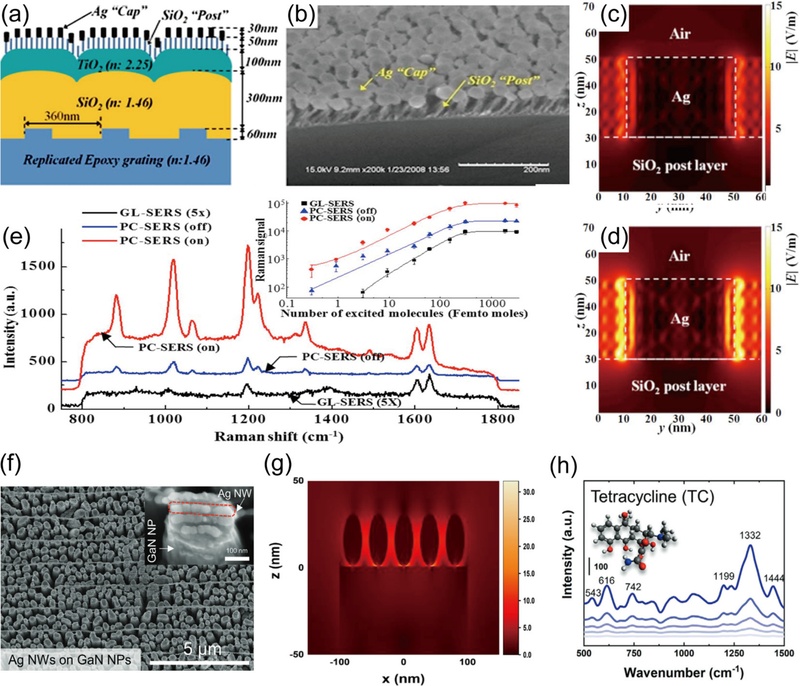
2-Dimensional PhC structured SERS. (a) Schematic of the PhC-SERS composition. (b) SEM image of the PhC-SERS. Cross-sectional electric field distribution of Ag-SiO2 nanorods on (c) GL-SERS and (d) PhC-SERS. (e) Comparison of Raman spectra of BPE obtained from GL-SERS and PhC-SERS substrates under on/off resonance conditions. Reproduced with permission from ref. [37]. Copyright (2008) AIP Publishing LLC. (f) SEM images of AgNW–GaN nanopillar plasmonic hybrid structures. (g) Cross-sectional electric field distribution of AgNWs on GaN nanopillars. (h) Raman spectrum of tetracycline showing the SERS effect of the GaN–Ag structure. Reproduced under the terms of the CC BY 4.0 license from ref. [38]. Copyright (2021) John Wiley and Sons.
For another case of a 2-D PhC structure applied to SERS, Lee et al. [38] demonstrated a GaN nanopillars-Ag nanowire (AgNW) plasmonic hybrid PhC structure. Fig. 4 (f) depicts SEM images of AgNWs formed on the GaN nanopillars. The nanopillars were randomly distributed with an average height of 2 μm and a diameter ranging from 200 to 300 nm. The cross-sectional electric field distribution of the GaN nanopillars-AgNWs revealed an extended electric field area through the PhC structure (Fig. 4 (g)). Accordingly, highly reproducible SERS signals of 10-14 M R6G on GaN nanopillars-AgNWs were observed. Moreover, the extended electric field area enabled the measurement of the biomolecule tetracycline, which has a relatively large size of 480.9 g/mol (Fig. 4 (h)). The research was progressive and proved that biomolecules considerably larger than the typical hot spot size can be measured through the electric field area expanded by the PhC structure.
In the case of 3-D PhC SERS, Jeong et al. [39] demonstrated a woodpile-structured SERS substrate. As shown in Fig. 5 (a), the 3-D woodpile PhC structure was composed of stacked AuNW arrays. The optimized stack number and the electric field distribution of the cross-section were investigated through FDTD simulation (Fig. 5 (b)). From the results, the 4 or 5-layer stacked woodpile structure had the maximum electric field enhancement (Fig. 5 (c)). To further verify the optimized structure experimentally, they fabricated and investigated 1 to 10-layer stacked woodpile structures (Fig. 5 (d)). Consistent with the results of FDTD simulation, it was revealed that the Raman enhancement for different number of stacked layers was saturated from 5 layers. According to the results, Park et al. [40] conducted biosensing research utilizing the 4-layer stacked woodpile PhC structure [40]. They functionalized the Au woodpile structure with carboxylic acid to measure the biomarkers of Alzheimer's disease, i.e., tau protein and amyloid β (Aβ). As shown in Fig. 5 (e), the Raman spectra of tau protein were measured even at 10-11 M with the woodpile SERS structure. Furthermore, they quantified the amount of tau protein by plotting the calibration curve for Raman intensity of the 1002 cm-1 peaks (Fig. 5 (f)). In addition, the Raman spectra of Aβ at 10-9 M was measured as the lowest concentration (Fig. 5 (g)). It was also quantified by the Raman intensity of the 1145 cm-1, 1270 cm-1, and 1380 cm-1 peaks of the Aβ (Fig. 5 (h)). Among the calibration curves, the blue line for the 1380 cm-1 peaks showed the most linear property. Consequently, it was proved possible to measure macromolecules such as proteins with the 3-D PhC SERS structure inducing the extended electric field area that occurs by the LSPR.
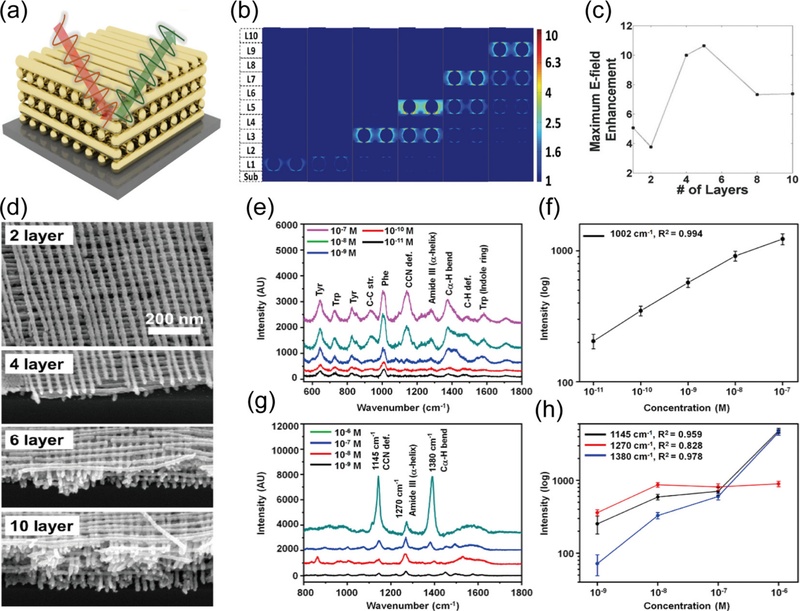
Three-dimensional PhC-structured SERS. (a) Schematic of a woodpile structure composed of AuNW arrays. (b) FDTD-simulated electric field distribution as a function of the number of stacked AuNW array layers. (c) Maximum electric field enhancement versus the number of layers. (d) SEM images of the stacked AuNW array layers. Reproduced with permission from ref. [39]. Copyright (2016) John Wiley & Sons. (e) Raman spectra of tau protein at various concentrations. (f) Calibration plot of the Raman intensity at the 1002 cm-1 peak in tau protein spectra as a function of concentration. (g) Raman spectra of amyloid β at various concentrations. (h) Calibration plots of the Raman intensities at the 1145 cm-1, 1270 cm-1, and 1380 cm-1 peaks in amyloid β spectra as a function of concentration. Reproduced with permission from ref. [40]. Copyright (2020) American Chemical Society.
Further advancing this concept, 3-D PhC SERS platforms were proposed. For example, Jeong et al. [39] introduced a structure based on vertically stacked gold nanowire arrays, and confirmed that a five-layer configuration maximized electric field enhancement through vertical coupling [39]. Building on this finding, Park et al. [40] demonstrated the utility of the structure in biosensing by detecting Alzheimer’s disease biomarkers such as tau protein and amyloid β at ultralow concentrations. More recently, Lee et al. [41] expanded the practicality of such architectures by developing a photolithography-free 3-D PhC SERS substrate with 80 nm-period nanopillars, highlighting the scalability and manufacturing flexibility of high-performance SERS designs.
3.3 Surface Functionalization
The presence of numerous biomolecules in the analyte matrix aside from the target analytes of interest pose challenges to use SERS as a biosensor [16]. For example, there are various biomolecules such as proteins and metabolites in body fluids [42]. The presence of background substances can pose a significant challenge in accurately identifying target analytes, obscuring the signals from the target analyte. Hence, an abundance of background substances represents a critical hurdle in the process of accurate identification. To address this issue, researchers are attempting to selectively attract the desired analytes near the hot spot by functionalizing the SERS substrate surface with materials such as polymers, aptamers and self-assembled monolayers (SAMs). Such strategic approaches can circumvent interference from the background biomolecules and increase the accuracy of detection [43].
Polymers are widely used due to their ability to interact with target analytes and attract biomolecules towards the hot spots [44]. Moreover, functionalized biocompatible polymers such as polyethylene glycol (PEG) [45], polyethylene imide (PEI) [46], and hyaluronic acid (HA) [47] stabilize the SERS substrate, allowing in vivo detection of biomolecules.
Son et al. [48] fabricated poly(diallyldimethylammonium chloride)-functionalized silver nanoshells (AgNS@PDDA) for real-time monitoring of multiple stress signaling molecules in living plants (Fig. 6 (a)). They first synthesized silver nanoshells (AgNS) with a bumpy surface to achieve strong Raman enhancement (Fig. 6 (b)). Cationic PDDA was then functionalized on the surface of the AgNS to improve biocompatibility and attract biomolecules to the AgNS surface. PDDA attracted plant stress-related molecules such as adenosine triphosphate (ATP), salicylic acid (SA), folic acid (FA), and indole acetic acid (IAA) through electrostatic interactions. The fingerprint spectral pattern of the analytes appeared in the SERS spectrum of both biomolecules and mixed compounds, demonstrating the multiplex detection performance of AgNS@PDDA (Fig. 6 (c)). The AgNS@PDDA was then infiltrated into the living plants and SERS spectra were acquired after wounding, cold stress (abiotic stress), and pathogen invasion (biotic stress). Distinct peaks corresponding to wound signaling molecules such as phytoalexin (1353 cm-1) and ATP (729 cm-1) were observed in the SERS spectra after wound stress (Fig. 6 (d)). Additionally, a prominent SERS peak of glutathione (643 cm-1) appeared after exposure to cold stress. The distinct SERS bands of SA and ATP, substances involved in resistance against pathogen invasion, were prominently detected after pathogen infection (Fig. 6 (e)).
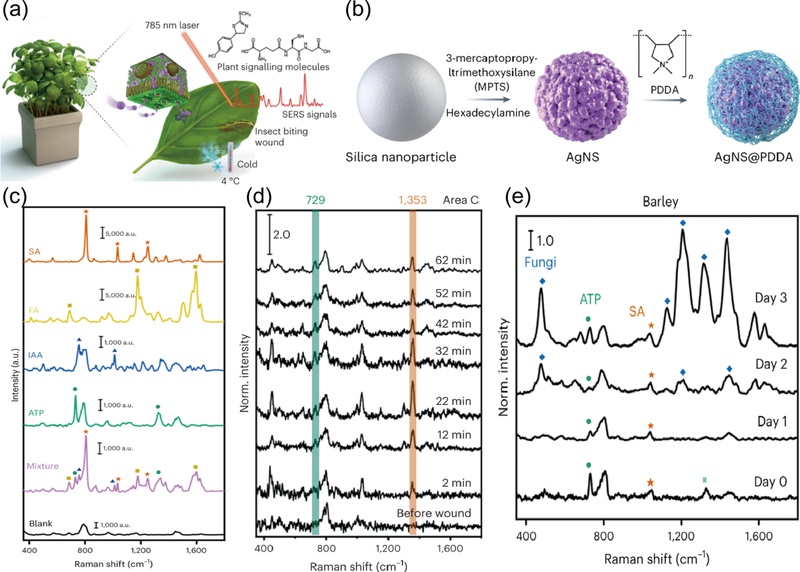
SERS sensor with polymer functionalization. (a) Schematic illustration of AgNS@PDDA for monitoring stress-related molecules in living plants. (b) Illustration of the AgNS@PDDA fabrication process. (c) SERS spectra of individual plant stress-related molecules and their mixture. (d) Real-time SERS spectra acquired from living plants after wound stress, and (e) after pathogen invasion. Reproduced with permission from ref. [48]. Copyright (2022) The Authors.
Choi et al. [45] detected diverse biomarkers expressed by cancer cells using Ag-Au hollow nanospheres encapsulated with a PEG layer (PEG-Ag-Au hollow). The PEG layer improved the biocompatibility and increased the stability of the SERS nanospheres. When the PEG-Ag-Au hollows were incubated with the cell, they showed considerably higher cell viability and proliferation compared to the pristine Ag-Au hollow. Jiang et al. [49] synthesized spiky gold nanoparticles (AuNPs) functionalized with HA and chitosan (CS) for imaging and visualization of cancer cells. HA enhanced the biocompatibility and water solubility of the spiky AuNPs. Moreover, it introduced targeting ability to the AuNPs because of affinity for the CD44 receptor on the membrane of the cancer cell.
Beyond polymers, additional strategies for surface functionalization have been proposed to enhance molecular specificity and analytical precision in SERS platforms. For instance, Cho et al. [50] employed short DNA aptamers to impart target-selective binding capability on metallic nanostructures, enabling the efficient discrimination of specific small molecules in chemically complex environments. Meanwhile, Kim et al. [51] introduced a range of SAMs with distinct chemical functionalities onto nanowire-based SERS substrates. This approach allowed for tunable analyte–surface interactions, thereby facilitating differentiation between metabolite profiles even within challenging biological matrices. When integrated with high-density 3-D structures such as woodpile architectures, these functionalization methods help maintain spectral clarity, while expanding detection capability to a broader class of biomolecules.
4. CONCLUSIONS
SERS has attracted significant attention for its potential in biosensing due to its exceptional sensitivity and molecular specificity. However, its practical use remains largely confined to controlled laboratory settings. To enable realworld applications, substantial efforts are underway to enhance sensing performance through structural and material innovation.
This review highlights recent advances in three key areas: the integration of ISs, the expansion of effective electromagnetic field regions, and the selective surface functionalization of SERS substrates. Together, these strategies have improved quantification reliability, enhanced detection sensitivity, and increased analyte specificity, positioning SERS as a highly promising biosensing platform.
Despite these advances, several challenges persist. IS strategies face issues such as irregular analyte distribution, hot spot overlap, and competition between analytes and standards. While waveguide and PhC structures offer extended field enhancement, limitations in fabrication and integration still hinder practical deployment. Accurate biomolecule detection is further complicated by background Raman signals and unwanted molecular interactions in complex biological environments. Achieving selective binding without spectral interference remains essential. Nevertheless, recent studies show that appropriate material selection and structural design can address these limitations. The adaptability of SERS platforms—enabled by their broad design flexibility—offers a promising path forward. With continued innovation, SERS is poised to become a powerful analytical tool in biosensing and beyond.
Acknowledgments
This research was supported by the KAIST Jang Young Sil Fellow Program.
References
-
A.I. Pérez-Jiménez, D. Lyu, Z. Lu, G. Liu, B. Ren, Surface-enhanced Raman spectroscopy: benefits, trade-offs and future developments, Chem. Sci. 11 (2020) 4563–4577.
[https://doi.org/10.1039/D0SC00809E]

-
Y. Liu, M. Kim, S.H. Cho, Y.S. Jung, Vertically aligned nanostructures for a reliable and ultrasensitive SERS-active platform: Fabrication and engineering strategies, Nano Today 37 (2021) 101063.
[https://doi.org/10.1016/j.nantod.2020.101063]

-
M. Fleischmann, P.J. Hendra, A.J. McQuillan, Raman spectra of pyridine adsorbed at a silver electrode, Chem. Phys. Lett. 26 (1974) 163–166.
[https://doi.org/10.1016/0009-2614(74)85388-1]

-
D.L. Jeanmaire, R.P. Van Duyne, Surface Raman spectroelectrochemistry: Part I. Heterocyclic, aromatic, and aliphatic amines adsorbed on the anodized silver electrode, J. Electroanal. Chem. Interf. Electrochem. 84 (1977) 1–20.
[https://doi.org/10.1016/0368-1874(77)80399-7]

-
Z. Wang, S. Zong, L. Wu, D. Zhu, Y. Cui, SERS-activated platforms for immunoassay: probes, encoding methods, and applications, Chem. Rev. 117 (2017) 7910–7963.
[https://doi.org/10.1021/acs.chemrev.7b00027]

-
M.F. Cardinal, E. Vander Ende, R.A. Hackler, M.O. McAnally, P.C. Stair, G.C. Schatz, et al., Expanding applications of SERS through versatile nanomaterials engineering, Chem. Soc. Rev. 46 (2017) 3886–3903.
[https://doi.org/10.1039/C7CS00207F]

-
E. Rho, M. Kim, S.H. Cho, B. Choi, H. Park, H. Jang, et al., Separation-free bacterial identification in arbitrary media via deep neural network-based SERS analysis, Biosens. Bioelectron. 202 (2022) 113991.
[https://doi.org/10.1016/j.bios.2022.113991]

-
S.-Y. Ding, J. Yi, J.-F. Li, B. Ren, D.-Y. Wu, R. Panneerselvam, et al., Nanostructure-based plasmon-enhanced Raman spectroscopy for surface analysis of materials, Nat. Rev. Mater. 1 (2016) 1–16.
[https://doi.org/10.1038/natrevmats.2016.21]

-
H.K. Lee, Y.H. Lee, C.S.L. Koh, G.C. Phan-Quang, X. Han, C. L. Lay, et al., Designing surface-enhanced Raman scattering (SERS) platforms beyond hotspot engineering: emerging opportunities in analyte manipulations and hybrid materials, Chem. Soc. Rev. 48 (2019) 731–756.
[https://doi.org/10.1039/C7CS00786H]

-
L.M. Almehmadi, S.M. Curley, N.A. Tokranova, S.A. Tenenbaum, I.K. Lednev, Surface enhanced Raman spectroscopy for single molecule protein detection, Sci. Rep. 9 (2019) 12356.
[https://doi.org/10.1038/s41598-019-48650-y]

-
X.X. Han, R.S. Rodriguez, C.L. Haynes, Y. Ozaki, B. Zhao, Surface-enhanced Raman spectroscopy, Nat. Rev. Methods Primers 1 (2021) 87.
[https://doi.org/10.1038/s43586-021-00083-6]

-
S. Unser, I. Bruzas, J. He, L. Sagle, Localized surface plasmon resonance biosensing: current challenges and approaches, Sensors 15 (2015) 15684–15716.
[https://doi.org/10.3390/s150715684]

-
N.H. Kim, W. Hwang, K. Baek, M.R. Rohman, J. Kim, H.W. Kim, et al., Smart SERS hot spots: single molecules can be positioned in a plasmonic nanojunction using host–guest chemistry, J. Am. Chem. Soc. 140 (2018) 4705–4711.
[https://doi.org/10.1021/jacs.8b01501]

-
L. Xia, M. Chen, X. Zhao, Z. Zhang, J. Xia, H. Xu, et al., Visualized method of chemical enhancement mechanism on SERS and TERS, J. Raman Spectrosc. 45 (2014) 533–540.
[https://doi.org/10.1002/jrs.4504]

-
M. Thomas, S. Mühlig, T. Deckert-Gaudig, C. Rockstuhl, V. Deckert, P. Marquetand, Distinguishing chemical and electromagnetic enhancement in surface-enhanced Raman spectra: The case of para-nitrothiophenol, J. Raman Spectrosc. 44 (2013) 1497–1505.
[https://doi.org/10.1002/jrs.4377]

-
C. Zong, M. Xu, L.-J. Xu, T. Wei, X. Ma, X.-S. Zheng, et al., Surface-enhanced Raman spectroscopy for bioanalysis: reliability and challenges, Chem. Rev. 118 (2018) 4946–4980.
[https://doi.org/10.1021/acs.chemrev.7b00668]

-
A. Bonifacio, S. Cervo, V. Sergo, Label-free surface-enhanced Raman spectroscopy of biofluids: fundamental aspects and diagnostic applications, Anal. Bioanal. Chem. 407 (2015) 8265–8277.
[https://doi.org/10.1007/s00216-015-8697-z]

-
Y. Wang, S. Schlücker, Rational design and synthesis of SERS labels, Analyst 138 (2013) 2224–2238.
[https://doi.org/10.1039/c3an36866a]

-
Y. Liu, H. Zhou, Z. Hu, G. Yu, D. Yang, J. Zhao, Label and label-free based surface-enhanced Raman scattering for pathogen bacteria detection: A review, Biosens. Bioelectron. 94 (2017) 131–140.
[https://doi.org/10.1016/j.bios.2017.02.032]

-
L.A. Lane, X. Qian, S. Nie, SERS nanoparticles in medicine: from label-free detection to spectroscopic tagging, Chem. Rev. 115 (2015) 10489–10529.
[https://doi.org/10.1021/acs.chemrev.5b00265]

-
X.-S. Zheng, I.J. Jahn, K. Weber, D. Cialla-May, J. Popp, Label-free SERS in biological and biomedical applications: recent progress, current challenges and opportunities, Spectrochim. Acta A Mol. Biomol. Spectrosc. 197 (2018) 56–77.
[https://doi.org/10.1016/j.saa.2018.01.063]

-
H.-Y. Chen, M.-H. Lin, C.-Y. Wang, Y.-M. Chang, S. Gwo, Large-scale hot spot engineering for quantitative SERS at the single-molecule scale, J. Am. Chem. Soc. 137 (2015) 13698–13705.
[https://doi.org/10.1021/jacs.5b09111]

-
G. Li, Q. Hao, M. Li, X. Zhao, W. Song, X. Fan, et al., Quantitative SERS analysis by employing janus nanoparticles with internal standards, Adv. Mater. Interfaces 10 (2023) 2202127.
[https://doi.org/10.1002/admi.202202127]

-
E. Hahm, M.G. Cha, E.J. Kang, X.-H. Pham, S.H. Lee, H.-M. Kim, et al., Multilayer Ag-embedded silica nanostructure as a surface-enhanced Raman scattering-based chemical sensor with dual-function internal standards, ACS Appl. Mater. Interfaces 10 (2018) 40748–40755.
[https://doi.org/10.1021/acsami.8b12640]

-
X. Wang, M. Xiao, Y. Zou, W. Lai, H. Pei, M.F. Alam, et al., Fractal SERS nanoprobes for multiplexed quantitative gene profiling, Biosens. Bioelectron. 156 (2020) 112130.
[https://doi.org/10.1016/j.bios.2020.112130]

-
R. Mei, Y. Wang, Q. Yu, Y. Yin, R. Zhao, L. Chen, Gold nanorod array-bridged internal-standard SERS tags: from ultrasensitivity to multifunctionality, ACS Appl. Mater. Interfaces 12 (2019) 2059–2066.
[https://doi.org/10.1021/acsami.9b18292]

-
X. Jiang, Z. Tan, L. Lin, J. He, C. He, B.D. Thackray, et al., Surface-enhanced Raman nanoprobes with embedded standards for quantitative cholesterol detection, Small Methods 2 (2018) 1800182.
[https://doi.org/10.1002/smtd.201800182]

-
H. Sun, X. Li, C. Gu, J. Zhang, G. Wei, T. Jiang, et al., Bioinspired surface-enhanced Raman scattering substrate with intrinsic Raman signal for the interactive SERS detection of pesticides residues, Spectrochim. Acta A Mol. Biomol. Spectrosc. 270 (2022) 120800.
[https://doi.org/10.1016/j.saa.2021.120800]

-
Z. Hao, S. Fu, H. Liu, H. Zhao, C. Gu, T. Jiang, Biomimetic SERS substrate with silicon-mediated internal standard: Improved sensing of environmental pollutants and nutrients, Spectrochim. Acta A Mol. Biomol. Spectrosc. 309 (2024) 123805.
[https://doi.org/10.1016/j.saa.2023.123805]

-
H. Tian, N. Zhang, L. Tong, J. Zhang, In situ quantitative graphene-based surface-enhanced Raman spectroscopy, Small Methods 1 (2017) 1700126.
[https://doi.org/10.1002/smtd.201700126]

-
Y. Ma, Y. Du, Y. Chen, C. Gu, T. Jiang, G. Wei, et al., Intrinsic Raman signal of polymer matrix induced quantitative multiphase SERS analysis based on stretched PDMS film with anchored Ag nanoparticles/Au nanowires, Chem. Eng. J. 381 (2020) 122710.
[https://doi.org/10.1016/j.cej.2019.122710]

-
J. Liu, Z. Hong, W. Yang, C. Liu, Z. Lu, L. Wu, et al., Bacteria inspired internal standard SERS substrate for quantitative detection, ACS Appl. Bio Mater. 4 (2020) 2009–2019.
[https://doi.org/10.1021/acsabm.0c00263]

-
H.-W. Cheng, H.-M. Tsai, Y.-L. Wang, Exploiting purine as an internal standard for SERS quantification of purine derivative molecules released by bacteria, Anal. Chem. 95 (2023) 16967–16975.
[https://doi.org/10.1021/acs.analchem.3c03259]

-
M. Fu, M. Mota, X. Xiao, A. Jacassi, N.A. Güsken, Y. Chen, et al., Near-unity Raman β-factor of surface-enhanced Raman scattering in a waveguide, Nat. Nanotechnol. 17 (2022) 1251–1257.
[https://doi.org/10.1038/s41565-022-01232-y]

-
A. Dhakal, P.C. Wuytens, F. Peyskens, K. Jans, N.L. Thomas, R. Baets, Nanophotonic waveguide enhanced Raman spectroscopy of biological submonolayers, ACS Photonics 3 (2016) 2141–2149.
[https://doi.org/10.1021/acsphotonics.6b00593]

-
S. V. Gaponenko, Introduction to nanophotonics, Cambridge University Press, Cambridge, 2010.
[https://doi.org/10.1017/CBO9780511750502]

-
S.-M. Kim, W. Zhang, B.T. Cunningham, Photonic crystals with SiO?–Ag “post-cap” nanostructure coatings for surface enhanced Raman spectroscopy, Appl. Phys. Lett. 93 (2008) 143117.
[https://doi.org/10.1063/1.2998695]

-
K.H. Lee, H. Jang, Y.S. Kim, C.H. Lee, S.H. Cho, M. Kim, H. Son, K.B. Bae, D.V. Dao, Y.S. Jung, Synergistic SERS enhancement in GaN–Ag hybrid system toward label-free and multiplexed detection of antibiotics in aqueous solutions, Adv. Sci. 8 (2021) 2100640.
[https://doi.org/10.1002/advs.202100640]

-
J.W. Jeong, M.M.P. Arnob, K.M. Baek, S.Y. Lee, W.C. Shih, Y.S. Jung, 3D cross-point plasmonic nanoarchitectures containing dense and regular hot spots for surface-enhanced Raman spectroscopy analysis, Adv. Mater. 28 (2016) 8695–8704.
[https://doi.org/10.1002/adma.201602603]

-
H.J. Park, S. Cho, M. Kim, Y.S. Jung, Carboxylic acid-functionalized, graphitic layer-coated three-dimensional SERS substrate for label-free analysis of Alzheimer’s disease biomarkers, Nano Lett. 20 (2020) 2576–2584.
[https://doi.org/10.1021/acs.nanolett.0c00048]

-
S. Lee, M. Ku, H. Lim, J. Hwang, J.M. Kim, H. Jang, et al., Realizing square-ordered nanopillars with a 0.1-tera-density through a superimposed masking strategy for advanced surface-enhanced Raman spectroscopy, ACS Appl. Mater. Interfaces 16 (2024) 69703–69712.
[https://doi.org/10.1021/acsami.4c15062]

-
B. Pelaz, G. Charron, C. Pfeiffer, Y. Zhao, J.M. De La Fuente, X.J. Liang, et al., Interfacing engineered nanoparticles with biological systems: anticipating adverse nano–bio interactions, Small 9 (2013) 1573–1584.
[https://doi.org/10.1002/smll.201201229]

-
T. Yaseen, H. Pu, D.-W. Sun, Functionalization techniques for improving SERS substrates and their applications in food safety evaluation: A review of recent research trends, Trends Food Sci. Technol. 72 (2018) 162–174.
[https://doi.org/10.1016/j.tifs.2017.12.012]

-
S. Fateixa, H.I. Nogueira, T. Trindade, SERS research applied to polymer based nanocomposites, In: G. M. do Nascimeto(Ed.), Raman Spectroscopy, 2018, pp. 91–126.
[https://doi.org/10.5772/intechopen.72680]

-
N. Choi, H. Dang, A. Das, M.S. Sim, I.Y. Chung, J. Choo, SERS biosensors for ultrasensitive detection of multiple biomarkers expressed in cancer cells, Biosens. Bioelectron. 164 (2020) 112326.
[https://doi.org/10.1016/j.bios.2020.112326]

-
F. Gu, S. Hu, Y. Wu, C. Wu, Y. Yang, B. Gu, et al., A SERS platform for rapid detection of drug resistance of non-Candida albicans using Fe?O?@PEI and triangular silver nanoplates, Int. J. Nanomed. 17 (2022) 3531–3541.
[https://doi.org/10.2147/IJN.S369591]

-
C. Hu, J. Shen, J. Yan, J. Zhong, W. Qin, R. Liu, et al., Highly narrow nanogap-containing Au@Au core–shell SERS nanoparticles: size-dependent Raman enhancement and applications in cancer cell imaging, Nanoscale 8 (2016) 2090–2096.
[https://doi.org/10.1039/C5NR06919J]

-
W.K. Son, Y.S. Choi, Y.W. Han, D.W. Shin, K. Min, J. Shin, et al., In vivo surface-enhanced Raman scattering nanosensor for the real-time monitoring of multiple stress signalling molecules in plants, Nat. Nanotechnol. 18 (2023) 205–216.
[https://doi.org/10.1038/s41565-022-01274-2]

-
R.T. Jiang, J.Q. Li, L.H. Li, Y.X. Lu, H.M. Zhang, M. Li, Engineering near-infrared plasmonic spiky gold nanostructures for highly efficient surface-enhanced Raman spectroscopy-guided cancer hyperthermia therapy, Adv. Funct. Mater. 34 (2024) 2307631.
[https://doi.org/10.1002/adfm.202307631]

-
S.H. Cho, K.M. Baek, H.J. Han, M. Kim, H. Park, Y.S. Jung, Selective, quantitative, and multiplexed surface-enhanced Raman spectroscopy using aptamer-functionalized monolithic plasmonic nanogrids derived from cross-point nano-welding, Adv. Funct. Mater. 30 (2020) 2000612.
[https://doi.org/10.1002/adfm.202000612]

-
M. Kim, S. Huh, H.J. Park, S.H. Cho, M.Y. Lee, S. Jo, et al., Surface-functionalized SERS platform for deep learning-assisted diagnosis of Alzheimer's disease, Biosens. Bioelectron. 251 (2024) 116128.
[https://doi.org/10.1016/j.bios.2024.116128]


Minjae Ku is currently a postdoctoral researcher in the Department of Materials Science and Engineering (MSE) at the Korea Advanced Institute of Science and Technology (KAIST). She earned a B.S. in MSE from the Ulsan National Institute of Science and Technology (UNIST) in 2018, an M.S. in MSE from Yonsei University in 2021, and a Ph.D. in MSE from KAIST in 2025. In 2021, she worked as a researcher at the Korea Research Institute of Bioscience and Biotechnology (KRIBB). Her research focuses on plasmonic nanomaterials and nanophotonics, with particular emphasis on metasurfaces, nanopatterning technologies, and light–matter interactions.
